Nonalcoholic steatohepatitis is associated with altered hepatic MicroRNA expression
- PMID: 19030170
- PMCID: PMC2717729
- DOI: 10.1002/hep.22569
Nonalcoholic steatohepatitis is associated with altered hepatic MicroRNA expression
Abstract
The expression of microRNA in nonalcoholic steatohepatitis (NASH) and their role in the genesis of NASH are not known. The aims of this study were to: (1) identify differentially expressed microRNAs in human NASH, (2) tabulate their potential targets, and (3) define the effect of a specific differentially expressed microRNA, miR-122, on its targets and compare these effects with the pattern of expression of these targets in human NASH. The expression of 474 human microRNAs was compared in subjects with the metabolic syndrome and NASH versus controls with normal liver histology. Differentially expressed microRNAs were identified by the muParaflo microRNA microarray assay and validated using quantitative real-time polymerase chain reaction (PCR). The effects of a specific differentially expressed miRNA (miR-122) on its predicted targets were assessed by silencing and overexpressing miR-122 in vitro. A total of 23 microRNAs were underexpressed or overexpressed. The predicted targets of these microRNAs are known to affect cell proliferation, protein translation, apoptosis, inflammation, oxidative stress, and metabolism. The miR-122 level was significantly decreased in subjects with NASH (63% by real-time PCR, P < 0.00001). Silencing miR-122 led to an initial increase in mRNA levels of these targets (P < 0.05 for all) followed by a decrease by 48 hours. This was accompanied by an increase in protein levels of these targets (P < 0.05 for all). Overexpression of miR-122 led to a significant decrease in protein levels of these targets.
Conclusions: NASH is associated with altered hepatic microRNA expression. Underexpression of miR-122 potentially contributes to altered lipid metabolism implicated in the pathogenesis of NASH.
Figures



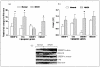



Comment in
-
Profiling microRNA expression: a snapshot of nonalcoholic steatohepatitis and a recording of its pathogenesis.Hepatology. 2009 Feb;49(2):706-7. doi: 10.1002/hep.22733. Hepatology. 2009. PMID: 19127512 No abstract available.
References
-
- Adams LA, Sanderson S, Lindor KD, Angulo P. The histological course of nonalcoholic fatty liver disease: a longitudinal study of 103 patients with sequential liver biopsies. J Hepatol. 2005;42:132–138. - PubMed
-
- Ambros V. The functions of animal microRNAs. Nature. 2004;431:350–355. - PubMed
-
- Nilsen TW. Mechanisms of microRNA-mediated gene regulation in animal cells. Trends Genet. 2007;23:243–249. - PubMed
-
- Lagos-Quintana M, Rauhut R, Yalcin A, Meyer J, Lendeckel W, Tuschl T. Identification of tissue-specific microRNAs from mouse. Curr Biol. 2002;12:735–739. - PubMed
-
- Houbaviy HB, Murray MF, Sharp PA. Embryonic stem cell-specific MicroRNAs. Dev Cell. 2003;5:351–358. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
