Identification of phylogenetically conserved sequence motifs in microRNA 5' flanking sites from C. elegans and C. briggsae
- PMID: 19036124
- PMCID: PMC2613404
- DOI: 10.1186/1471-2199-9-105
Identification of phylogenetically conserved sequence motifs in microRNA 5' flanking sites from C. elegans and C. briggsae
Abstract
Background: MicroRNAs (miRNAs) are small, noncoding RNA molecules that act as post-transcriptional regulators of gene expression. Studies concerning transcriptional regulation of miRNAs have so far concentrated on those located within the intergenic region of the genome and the search for putative promoters, thus leaving open the question of the existence of possible regulatory elements common to all miRNAs including those located in introns of protein coding genes.
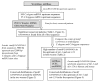
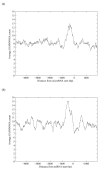
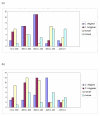
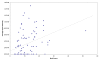
Results: In this study, we initially searched for motifs occurring in the area 1000 bp upstream from all miRNAs independent of their genomic location. We discovered a previously unknown sequence motif GANNNNGA that displayed a conserved distribution in the nematode worms Caenorhabditis elegans and Caenorhabditis briggsae. This motif had a peak occurrence at 500 bp upstream, with a sharp drop-off toward the miRNA start site. Further analysis indicated that this motif was locally restricted and not enriched 1000-5000 bp upstream or 0-2000 bp downstream of the miRNA start site. In addition, this motif was observed to be most abundant in the upstream sequences of two important miRNAs, mir-1 and mir-124. This abundance was also conserved in phylogenetically distant species including human and mouse.
Conclusion: The results show that the motif GANNNNGA is conserved close to miRNA precursor start sites, suggesting that it may be involved in miRNA sequence recognition or regulation. This data provides important knowledge for the identification and computational prediction of miRNA sequences.
Figures

Similar articles
-
Comparative genomic analysis of upstream miRNA regulatory motifs in Caenorhabditis.RNA. 2016 Jul;22(7):968-78. doi: 10.1261/rna.055392.115. Epub 2016 May 2. RNA. 2016. PMID: 27140965 Free PMC article.
-
Patterns of flanking sequence conservation and a characteristic upstream motif for microRNA gene identification.RNA. 2004 Sep;10(9):1309-22. doi: 10.1261/rna.5206304. RNA. 2004. PMID: 15317971 Free PMC article.
-
Repertoire and evolution of miRNA genes in four divergent nematode species.Genome Res. 2009 Nov;19(11):2064-74. doi: 10.1101/gr.093781.109. Epub 2009 Sep 15. Genome Res. 2009. PMID: 19755563 Free PMC article.
-
The draft genome sequence of the nematode Caenorhabditis briggsae, a companion to C. elegans.Genome Biol. 2003;4(12):238. doi: 10.1186/gb-2003-4-12-238. Epub 2003 Nov 18. Genome Biol. 2003. PMID: 14659008 Free PMC article. Review.
-
MicroRNAs: small RNAs with big effects.Transplantation. 2010 Jul 27;90(2):105-12. doi: 10.1097/TP.0b013e3181e913c2. Transplantation. 2010. PMID: 20574417 Free PMC article. Review.
Cited by
-
Transcriptional control of microRNA expression in C. elegans: promoting better understanding.RNA Biol. 2009 Jan-Mar;6(1):49-53. doi: 10.4161/rna.6.1.7574. Epub 2009 Jan 9. RNA Biol. 2009. PMID: 19106630 Free PMC article. Review.
-
Comparative genomic analysis of upstream miRNA regulatory motifs in Caenorhabditis.RNA. 2016 Jul;22(7):968-78. doi: 10.1261/rna.055392.115. Epub 2016 May 2. RNA. 2016. PMID: 27140965 Free PMC article.
-
Characterization of rubber tree microRNA in phytohormone response using large genomic DNA libraries, promoter sequence and gene expression analysis.Mol Genet Genomics. 2014 Oct;289(5):921-33. doi: 10.1007/s00438-014-0862-0. Epub 2014 May 26. Mol Genet Genomics. 2014. PMID: 24859131
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

