Genetics meets metabolomics: a genome-wide association study of metabolite profiles in human serum
- PMID: 19043545
- PMCID: PMC2581785
- DOI: 10.1371/journal.pgen.1000282
Genetics meets metabolomics: a genome-wide association study of metabolite profiles in human serum
Abstract
The rapidly evolving field of metabolomics aims at a comprehensive measurement of ideally all endogenous metabolites in a cell or body fluid. It thereby provides a functional readout of the physiological state of the human body. Genetic variants that associate with changes in the homeostasis of key lipids, carbohydrates, or amino acids are not only expected to display much larger effect sizes due to their direct involvement in metabolite conversion modification, but should also provide access to the biochemical context of such variations, in particular when enzyme coding genes are concerned. To test this hypothesis, we conducted what is, to the best of our knowledge, the first GWA study with metabolomics based on the quantitative measurement of 363 metabolites in serum of 284 male participants of the KORA study. We found associations of frequent single nucleotide polymorphisms (SNPs) with considerable differences in the metabolic homeostasis of the human body, explaining up to 12% of the observed variance. Using ratios of certain metabolite concentrations as a proxy for enzymatic activity, up to 28% of the variance can be explained (p-values 10(-16) to 10(-21)). We identified four genetic variants in genes coding for enzymes (FADS1, LIPC, SCAD, MCAD) where the corresponding metabolic phenotype (metabotype) clearly matches the biochemical pathways in which these enzymes are active. Our results suggest that common genetic polymorphisms induce major differentiations in the metabolic make-up of the human population. This may lead to a novel approach to personalized health care based on a combination of genotyping and metabolic characterization. These genetically determined metabotypes may subscribe the risk for a certain medical phenotype, the response to a given drug treatment, or the reaction to a nutritional intervention or environmental challenge.
Conflict of interest statement
KMW is an employee of Biocrates Life Sciences AG. This private company offers products and services in the field of targeted quantitative metabolomics research. The other authors have no competing interests to declare.
Figures

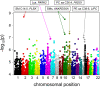


References
-
- McCarthy MI, Abecasis GR, Cardon LR, Goldstein DB, Little J, et al. Genome-wide association studies for complex traits: consensus, uncertainty and challenges. Nat Rev Genet. 2008;9:356–369. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

