Reference alignment of SNP microarray signals for copy number analysis of tumors
- PMID: 19052058
- PMCID: PMC2639073
- DOI: 10.1093/bioinformatics/btn624
Reference alignment of SNP microarray signals for copy number analysis of tumors
Abstract
A new procedure to align single nucleotide polymorphism (SNP) microarray signals for copy number analysis is proposed. For each individual array, this reference alignment procedure (RAP) uses a set of selected markers as internal references to direct the signal alignment. RAP aligns the signals so that each array has a similar signal distribution among its reference markers. An accompanying reference selection algorithm (RSA) uses genotype calls and initial signal intensities to choose two-copy markers as the internal references for each array. After RSA and RAP are applied, each array has a similar distribution of signals of two-copy markers so that across-array signal comparisons are biologically meaningful. An upper bound for a statistical metric of signal misalignment is derived and provides a theoretical basis to choose RSA-RAP over other alignment procedures for copy number analysis of cancers. In our study of acute lymphoblastic leukemia, RSA-RAP gives copy number analysis results that show substantially better concordance with cytogenetics than do two other alignment procedures.
Availability: Documented R code is freely available from www.stjuderesearch.org/depts/biostats/refnorm.
Figures
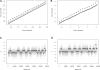

References
-
- Bolstad BM, et al. A comparison of normalization methods for high density oligonucleotide array based on variance and bias. Bioinformatics. 2003;19:185–193. - PubMed
-
- Mullighan CG, et al. Genes regulating B cell development are mutated in acute lymphoid leukaemia. Nature. 2007;446:758–764.
-
- Pounds S, Cheng C. Statistical development and evaluation of gene expression data filters. J. Comput. Biol. 2005;12:482–495. - PubMed
-
- Raimondi SC, et al. Cytogenetics as a diagnostic aid for childhood hematologic disorders: conventional cytogenetic techniques, fluorescence in situ hybridization, and comparative genomic hybridization. In: Hanausek M, Walaszek Z, editors. Tumor Marker Protocols. Methods Molecular Medicine. Totowa, NJ: Humana Press; 1998. pp. 209–227. 1998.

