ADAR1 is essential for the maintenance of hematopoiesis and suppression of interferon signaling
- PMID: 19060901
- PMCID: PMC2701568
- DOI: 10.1038/ni.1680
ADAR1 is essential for the maintenance of hematopoiesis and suppression of interferon signaling
Erratum in
- Nat Immunol. 2009 May;10(5):551
Abstract
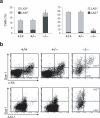
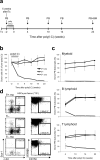
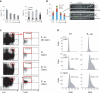
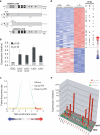
The deaminase ADAR1 edits adenosines in nuclear transcripts of nervous tissue and is required in the fetal liver of the developing mouse embryo. Here we show by inducible gene disruption in mice that ADAR1 is essential for maintenance of both fetal and adult hematopoietic stem cells. Loss of ADAR1 in hematopoietic stem cells led to global upregulation of type I and II interferon-inducible transcripts and rapid apoptosis. Our findings identify ADAR1 as an essential regulator of hematopoietic stem cell maintenance and suppressor of interferon signaling that may protect organisms from the deleterious effects of interferon activation associated with many pathological processes, including chronic inflammation, autoimmune disorders and cancer.
Figures
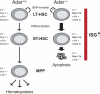
Comment in
-
A new function for the RNA-editing enzyme ADAR1.Nat Immunol. 2009 Jan;10(1):16-8. doi: 10.1038/ni0109-16. Nat Immunol. 2009. PMID: 19088736 Free PMC article.
References
-
- Levanon EY, et al. Systematic identification of abundant A-to-I editing sites in the human transcriptome. Nature biotechnology. 2004;22:1001–1005. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

