Genome-wide expression profiling reveals distinct clusters of transcriptional regulation during bovine preimplantation development in vivo
- PMID: 19064908
- PMCID: PMC2604955
- DOI: 10.1073/pnas.0805616105
Genome-wide expression profiling reveals distinct clusters of transcriptional regulation during bovine preimplantation development in vivo
Erratum in
- Proc Natl Acad Sci U S A. 2009 Feb 3;106(5):1679
Abstract
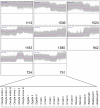
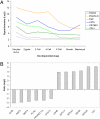
Bovine embryos can be generated by in vitro fertilization or somatic nuclear transfer; however, these differ from their in vivo counterparts in many aspects and exhibit a higher proportion of developmental abnormalities. Here, we determined for the first time the transcriptomes of bovine metaphase II oocytes and all stages of preimplantation embryos developing in vivo up to the blastocyst using the Affymetrix GeneChip Bovine Genome Array which examines approximately 23,000 transcripts. The data show that bovine oocytes and embryos transcribed a significantly higher number of genes than somatic cells. Several hundred genes were transcribed well before the 8-cell stage, at which the major activation of the bovine genome expression occurs. Importantly, stage-specific expression patterns in 2-cell, 4-cell, and 8-cell stages, and in morulae and blastocysts, were detected, indicating dynamic changes in the embryonic transcriptome and in groups of transiently active genes. Pathway analysis revealed >120 biochemical pathways that are operative in early preimplantation bovine development. Significant differences were observed between the mRNA expression profiles of in vivo and in vitro matured oocytes, highlighting the need to include in vivo derived oocytes/embryos in studies evaluating assisted reproductive techniques. This study provides the first comprehensive analysis of gene expression and transcriptome dynamics of in vivo developing bovine embryos and will serve as a basis for improving assisted reproductive technology.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
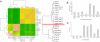
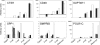
Similar articles
-
Conserved molecular portraits of bovine and human blastocysts as a consequence of the transition from maternal to embryonic control of gene expression.Physiol Genomics. 2007 Oct 22;31(2):315-27. doi: 10.1152/physiolgenomics.00041.2007. Epub 2007 Jun 26. Physiol Genomics. 2007. PMID: 17595343
-
Expression of RNA from developmentally important genes in preimplantation bovine embryos produced in TCM supplemented with BSA.J Reprod Fertil. 1998 Mar;112(2):387-98. doi: 10.1530/jrf.0.1120387. J Reprod Fertil. 1998. PMID: 9640278
-
Transcriptional profiles of bovine in vivo pre-implantation development.BMC Genomics. 2014 Sep 4;15(1):756. doi: 10.1186/1471-2164-15-756. BMC Genomics. 2014. PMID: 25185836 Free PMC article.
-
Gene expression analysis and in vitro production procedures for bovine preimplantation embryos: Past highlights, present concepts and future prospects.Reprod Domest Anim. 2018 Sep;53 Suppl 2:14-19. doi: 10.1111/rda.13260. Reprod Domest Anim. 2018. PMID: 30238652 Review.
-
Genome activation in bovine embryos: review of the literature and new insights from RNA sequencing experiments.Anim Reprod Sci. 2014 Sep;149(1-2):46-58. doi: 10.1016/j.anireprosci.2014.05.016. Epub 2014 Jun 6. Anim Reprod Sci. 2014. PMID: 24975847 Review.
Cited by
-
Histone modifications and mRNA expression in the inner cell mass and trophectoderm of bovine blastocysts.Epigenetics. 2013 Mar;8(3):281-9. doi: 10.4161/epi.23899. Epub 2013 Feb 13. Epigenetics. 2013. PMID: 23406883 Free PMC article.
-
A developmentally programmed splicing failure contributes to DNA damage response attenuation during mammalian zygotic genome activation.Sci Adv. 2022 Apr 15;8(15):eabn4935. doi: 10.1126/sciadv.abn4935. Epub 2022 Apr 13. Sci Adv. 2022. PMID: 35417229 Free PMC article.
-
Temporal patterns of gene regulation and upstream regulators contributing to major developmental transitions during Rhesus macaque preimplantation development.Mol Hum Reprod. 2019 Mar 1;25(3):111-123. doi: 10.1093/molehr/gaz001. Mol Hum Reprod. 2019. PMID: 30698740 Free PMC article.
-
Sequential analysis of global gene expression profiles in immature and in vitro matured bovine oocytes: potential molecular markers of oocyte maturation.BMC Genomics. 2011 Mar 16;12:151. doi: 10.1186/1471-2164-12-151. BMC Genomics. 2011. PMID: 21410957 Free PMC article.
-
Deregulation of oxidative phosphorylation pathways in embryos derived in vitro from prepubertal and pubertal heifers based on whole-transcriptome sequencing.BMC Genomics. 2024 Jun 24;25(1):632. doi: 10.1186/s12864-024-10532-7. BMC Genomics. 2024. PMID: 38914933 Free PMC article.
References
-
- Schultz RM. Regulation of zygotic gene activation in the mouse. Bioessays. 1993;15:531–538. - PubMed
-
- Braude P, Bolton V, Moore S. Human gene expression first occurs between the four- and eight-cell stages of preimplantation development. Nature. 1988;332:459–461. - PubMed
-
- Telford NA, Watson AJ, Schultz GA. Transition from maternal to embryonic control in early development: A comparison of several species. Mol Reprod Dev. 1990;26:90–100. - PubMed
-
- Kues WA, Niemann H. The contribution of farm animals to human health. Trends Biotechnol. 2004;22:286–294. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

