Analysis of cryo-electron microscopy images does not support the existence of 30-nm chromatin fibers in mitotic chromosomes in situ
- PMID: 19064912
- PMCID: PMC2604964
- DOI: 10.1073/pnas.0810057105
Analysis of cryo-electron microscopy images does not support the existence of 30-nm chromatin fibers in mitotic chromosomes in situ
Abstract
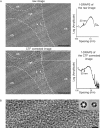
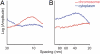
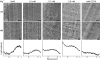
Although the formation of 30-nm chromatin fibers is thought to be the most basic event of chromatin compaction, it remains controversial because high-resolution imaging of chromatin in living eukaryotic cells had not been possible until now. Cryo-electron microscopy of vitreous sections is a relatively new technique, which enables direct high-resolution observation of the cell structures in a close-to-native state. We used cryo-electron microscopy and image processing to further investigate the presence of 30-nm chromatin fibers in human mitotic chromosomes. HeLa S3 cells were vitrified by high-pressure freezing, thin-sectioned, and then imaged under the cryo-electron microscope without any further chemical treatment or staining. For an unambiguous interpretation of the images, the effects of the contrast transfer function were computationally corrected. The mitotic chromosomes of the HeLa S3 cells appeared as compact structures with a homogeneous grainy texture, in which there were no visible 30-nm fibers. Power spectra of the chromosome images also gave no indication of 30-nm chromatin folding. These results, together with our observations of the effects of chromosome swelling, strongly suggest that, within the bulk of compact metaphase chromosomes, the nucleosomal fiber does not undergo 30-nm folding, but exists in a highly disordered and interdigitated state, which is, on the local scale, comparable with a polymer melt.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Comment in
-
A silent revolution in chromosome biology.Nat Rev Mol Cell Biol. 2014 Jul;15(7):431. doi: 10.1038/nrm3817. Epub 2014 May 29. Nat Rev Mol Cell Biol. 2014. PMID: 24871800 No abstract available.
References
-
- Becker WM, Reece JB, Poenie MF. In: The World of the Cell. Becker WM, editor. Menlo Park, CA: Benjamin-Cummings Publishing Company; 1996. pp. 434–435.
-
- Kornberg RD. Chromatin structure: A repeating unit of histones and DNA. Science. 1974;184:868–871. - PubMed
-
- Oudet P, Gross-Bellard M, Chambon P. Electron microscopic and biochemical evidence that chromatin structure is a repeating unit. Cell. 1975;4:281–300. - PubMed
-
- Kornberg RD, Lorch Y. Twenty-five years of the nucleosome, fundamental particle of the eukaryote chromosome. Cell. 1999;98:285–294. - PubMed
-
- Swedlow JR, Hirano T. The making of the mitotic chromosome: Modern insights into classical questions. Mol Cell. 2003;11:557–569. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

