Combinatorial regulation of endothelial gene expression by ets and forkhead transcription factors
- PMID: 19070576
- PMCID: PMC2782666
- DOI: 10.1016/j.cell.2008.10.049
Combinatorial regulation of endothelial gene expression by ets and forkhead transcription factors
Abstract
Vascular development begins when mesodermal cells differentiate into endothelial cells, which then form primitive vessels. It has been hypothesized that endothelial-specific gene expression may be regulated combinatorially, but the transcriptional mechanisms governing specificity in vascular gene expression remain incompletely understood. Here, we identify a 44 bp transcriptional enhancer that is sufficient to direct expression specifically and exclusively to the developing vascular endothelium. This enhancer is regulated by a composite cis-acting element, the FOX:ETS motif, which is bound and synergistically activated by Forkhead and Ets transcription factors. We demonstrate that coexpression of the Forkhead protein FoxC2 and the Ets protein Etv2 induces ectopic expression of vascular genes in Xenopus embryos, and that combinatorial knockdown of the orthologous genes in zebrafish embryos disrupts vascular development. Finally, we show that FOX:ETS motifs are present in many known endothelial-specific enhancers and that this motif is an efficient predictor of endothelial enhancers in the human genome.
Figures




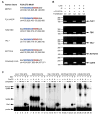


References
-
- Andreoli CM, Miller JW. Anti-vascular endothelial growth factor therapy for ocular neovascular disease. Curr Opin Ophthalmol. 2007;18:502–508. - PubMed
-
- Baron MH. Embryonic origins of mammalian hematopoiesis. Exp Hematol. 2003;31:1160–1169. - PubMed
-
- Carlsson P, Mahlapuu M. Forkhead transcription factors: key players in development and metabolism. Dev Biol. 2002;250:1–23. - PubMed
-
- Carmeliet P, Jain RK. Angiogenesis in cancer and other diseases. Nature. 2000;407:249–257. - PubMed
-
- Cleaver O, Tonissen KF, Saha MS, Krieg PA. Neovascularization of the Xenopus embryo. Dev Dyn. 1997;210:66–77. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

