Sgt1 dimerization is required for yeast kinetochore assembly
- PMID: 19073600
- PMCID: PMC2635038
- DOI: 10.1074/jbc.M806281200
Sgt1 dimerization is required for yeast kinetochore assembly
Abstract
The kinetochore, which consists of DNA sequence elements and structural proteins, is essential for high-fidelity chromosome transmission during cell division. In budding yeast, Sgt1 and Hsp90 help assemble the core kinetochore complex CBF3 by activating the CBF3 components Skp1 and Ctf13. In this study, we show that Sgt1 forms homodimers by performing in vitro and in vivo immunoprecipitation and analytical ultracentrifugation analyses. Analyses of the dimerization of Sgt1 deletion proteins showed that the Skp1-binding domain (amino acids 1-211) contains the Sgt1 homodimerization domain. Also, the Sgt1 mutant proteins that were unable to dimerize also did not bind Skp1, suggesting that Sgt1 dimerization is important for Sgt1-Skp1 binding. Restoring dimerization activity of a dimerization-deficient sgt1 mutant (sgt1-L31P) by using the CENP-B (centromere protein-B) dimerization domain suppressed the temperature sensitivity, the benomyl sensitivity, and the chromosome missegregation phenotype of sgt1-L31P. These results strongly suggest that Sgt1 dimerization is required for kinetochore assembly.
Figures



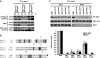

Similar articles
-
Sgt1 associates with Hsp90: an initial step of assembly of the core kinetochore complex.Mol Cell Biol. 2004 Sep;24(18):8069-79. doi: 10.1128/MCB.24.18.8069-8079.2004. Mol Cell Biol. 2004. PMID: 15340069 Free PMC article.
-
Sgt1 dimerization is negatively regulated by protein kinase CK2-mediated phosphorylation at Ser361.J Biol Chem. 2009 Jul 10;284(28):18692-8. doi: 10.1074/jbc.M109.012732. Epub 2009 Apr 27. J Biol Chem. 2009. PMID: 19398558 Free PMC article.
-
The crystal structure of the Sgt1-Skp1 complex: the link between Hsp90 and both SCF E3 ubiquitin ligases and kinetochores.Sci Rep. 2017 Jan 31;7:41626. doi: 10.1038/srep41626. Sci Rep. 2017. PMID: 28139700 Free PMC article.
-
Structures and functions of yeast kinetochore complexes.Annu Rev Biochem. 2007;76:563-91. doi: 10.1146/annurev.biochem.76.052705.160607. Annu Rev Biochem. 2007. PMID: 17362199 Review.
-
Chromatin assembly: the kinetochore connection.Curr Biol. 2002 Apr 2;12(7):R256-8. doi: 10.1016/s0960-9822(02)00786-8. Curr Biol. 2002. PMID: 11937044 Review.
Cited by
-
A designed point mutant in Fis1 disrupts dimerization and mitochondrial fission.J Mol Biol. 2012 Oct 19;423(2):143-58. doi: 10.1016/j.jmb.2012.06.042. Epub 2012 Jul 9. J Mol Biol. 2012. PMID: 22789569 Free PMC article.
-
Origin of a folded repeat protein from an intrinsically disordered ancestor.Elife. 2016 Sep 13;5:e16761. doi: 10.7554/eLife.16761. Elife. 2016. PMID: 27623012 Free PMC article.
-
Interplay between the phosphatase PHLPP1 and E3 ligase RNF41 stimulates proper kinetochore assembly via the outer-kinetochore protein SGT1.J Biol Chem. 2017 Aug 25;292(34):13947-13958. doi: 10.1074/jbc.M117.782896. Epub 2017 Jul 10. J Biol Chem. 2017. PMID: 28696259 Free PMC article.
-
Genes encoding Cher-TPR fusion proteins are predominantly found in gene clusters encoding chemosensory pathways with alternative cellular functions.PLoS One. 2012;7(9):e45810. doi: 10.1371/journal.pone.0045810. Epub 2012 Sep 20. PLoS One. 2012. PMID: 23029255 Free PMC article.
-
Mechanisms of Hsp90 regulation.Biochem J. 2016 Aug 15;473(16):2439-52. doi: 10.1042/BCJ20160005. Biochem J. 2016. PMID: 27515256 Free PMC article. Review.
References
-
- Cheeseman, I. M., and Desai, A. (2008) Nat. Rev. Mol. Cell Biol. 9 33–46 - PubMed
-
- Westermann, S., Drubin, D. G., and Barnes, G. (2007) Annu. Rev. Biochem. 76 563–591 - PubMed
-
- Sudakin, V., and Yen, T. J. (2007) BioDrugs 21 225–233 - PubMed
-
- Weaver, B. A., Silk, A. D., Montagna, C., Verdier-Pinard, P., and Cleveland, D. W. (2007) Cancer Cell 11 25–36 - PubMed
-
- Fitzgerald-Hayes, M., Clarke, L., and Carbon, J. (1982) Cell 29 235–244 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

