Alzheimer disease Abeta production in the absence of S-palmitoylation-dependent targeting of BACE1 to lipid rafts
- PMID: 19074428
- PMCID: PMC2635050
- DOI: 10.1074/jbc.M808920200
Alzheimer disease Abeta production in the absence of S-palmitoylation-dependent targeting of BACE1 to lipid rafts
Abstract
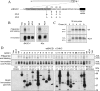
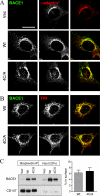
Alzheimer disease beta-amyloid (Abeta) peptides are generated via sequential proteolysis of amyloid precursor protein (APP) by BACE1 and gamma-secretase. A subset of BACE1 localizes to cholesterol-rich membrane microdomains, termed lipid rafts. BACE1 processing in raft microdomains of cultured cells and neurons was characterized in previous studies by disrupting the integrity of lipid rafts by cholesterol depletion. These studies found either inhibition or elevation of Abeta production depending on the extent of cholesterol depletion, generating controversy. The intricate interplay between cholesterol levels, APP trafficking, and BACE1 processing is not clearly understood because cholesterol depletion has pleiotropic effects on Golgi morphology, vesicular trafficking, and membrane bulk fluidity. In this study, we used an alternate strategy to explore the function of BACE1 in membrane microdomains without altering the cellular cholesterol level. We demonstrate that BACE1 undergoes S-palmitoylation at four Cys residues at the junction of transmembrane and cytosolic domains, and Ala substitution at these four residues is sufficient to displace BACE1 from lipid rafts. Analysis of wild type and mutant BACE1 expressed in BACE1 null fibroblasts and neuroblastoma cells revealed that S-palmitoylation neither contributes to protein stability nor subcellular localization of BACE1. Surprisingly, non-raft localization of palmitoylation-deficient BACE1 did not have discernible influence on BACE1 processing of APP or secretion of Abeta. These results indicate that post-translational S-palmitoylation of BACE1 is not required for APP processing, and that BACE1 can efficiently cleave APP in both raft and non-raft microdomains.
Figures
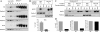



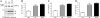
References
-
- Vassar, R., Bennett, B. D., Babu-Khan, S., Kahn, S., Mendiaz, E. A., Denis, P., Teplow, D. B., Ross, S., Amarante, P., Loeloff, R., Luo, Y., Fisher, S., Fuller, J., Edenson, S., Lile, J., Jarosinski, M. A., Biere, A. L., Curran, E., Burgess, T., Louis, J. C., Collins, F., Treanor, J., Rogers, G., and Citron, M. (1999) Science 286 735-741 - PubMed
-
- Sinha, S., Anderson, J. P., Barbour, R., Basi, G. S., Caccavello, R., Davis, D., Doan, M., Dovey, H. F., Frigon, N., Hong, J., Jacobson-Croak, K., Jewett, N., Keim, P., Knops, J., Lieberburg, I., Power, M., Tan, H., Tatsuno, G., Tung, J., Schenk, D., Seubert, P., Suomensaari, S. M., Wang, S., Walker, D., Zhao, J., McConlogue, L., and John, V. (1999) Nature 402 537-540 - PubMed
-
- Yan, R., Bienkowski, M. J., Shuck, M. E., Miao, H., Tory, M. C., Pauley, A. M., Brashier, J. R., Stratman, N. C., Mathews, W. R., Buhl, A. E., Carter, D. B., Tomasselli, A. G., Parodi, L. A., Heinrikson, R. L., and Gurney, M. E. (1999) Nature 402 533-537 - PubMed
-
- Cai, H., Wang, Y., McCarthy, D., Wen, H., Borchelt, D. R., Price, D. L., and Wong, P. C. (2001) Nat. Neurosci. 4 233-234 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

