Duplication and gene conversion in the Drosophila melanogaster genome
- PMID: 19079581
- PMCID: PMC2588116
- DOI: 10.1371/journal.pgen.1000305
Duplication and gene conversion in the Drosophila melanogaster genome
Abstract
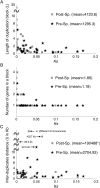
Using the genomic sequences of Drosophila melanogaster subgroup, the pattern of gene duplications was investigated with special attention to interlocus gene conversion. Our fine-scale analysis with careful visual inspections enabled accurate identification of a number of duplicated blocks (genomic regions). The orthologous parts of those duplicated blocks were also identified in the D. simulans and D. sechellia genomes, by which we were able to clearly classify the duplicated blocks into post- and pre-speciation blocks. We found 31 post-speciation duplicated genes, from which the rate of gene duplication (from one copy to two copies) is estimated to be 1.0 x 10(-9) per single-copy gene per year. The role of interlocus gene conversion was observed in several respects in our data: (1) synonymous divergence between a duplicated pair is overall very low. Consequently, the gene duplication rate would be seriously overestimated by counting duplicated genes with low divergence; (2) the sizes of young duplicated blocks are generally large. We postulate that the degeneration of gene conversion around the edges could explain the shrinkage of "identifiable" duplicated regions; and (3) elevated paralogous divergence is observed around the edges in many duplicated blocks, supporting our gene conversion-degeneration model. Our analysis demonstrated that gene conversion between duplicated regions is a common and genome-wide phenomenon in the Drosophila genomes, and that its role should be especially significant in the early stages of duplicated genes. Based on a population genetic prediction, we applied a new genome-scan method to test for signatures of selection for neofunctionalization and found a strong signature in a pair of transporter genes.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

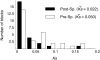

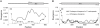

Similar articles
-
Population genetics of duplicated alternatively spliced exons of the Dscam gene in Daphnia and Drosophila.PLoS One. 2011;6(12):e27947. doi: 10.1371/journal.pone.0027947. Epub 2011 Dec 12. PLoS One. 2011. PMID: 22174757 Free PMC article.
-
Evolutionary dynamics of recently duplicated genes: Selective constraints on diverging paralogs in the Drosophila pseudoobscura genome.J Mol Evol. 2009 Jul;69(1):81-93. doi: 10.1007/s00239-009-9254-1. Epub 2009 Jun 18. J Mol Evol. 2009. PMID: 19536449
-
Drosophila duplication hotspots are associated with late-replicating regions of the genome.PLoS Genet. 2011 Nov;7(11):e1002340. doi: 10.1371/journal.pgen.1002340. Epub 2011 Nov 3. PLoS Genet. 2011. PMID: 22072977 Free PMC article.
-
Birth and death of duplicated genes in completely sequenced eukaryotes.Trends Genet. 2001 May;17(5):237-9. doi: 10.1016/s0168-9525(01)02243-0. Trends Genet. 2001. PMID: 11335019 Review.
-
An Overview of Duplicated Gene Detection Methods: Why the Duplication Mechanism Has to Be Accounted for in Their Choice.Genes (Basel). 2020 Sep 4;11(9):1046. doi: 10.3390/genes11091046. Genes (Basel). 2020. PMID: 32899740 Free PMC article. Review.
Cited by
-
Protein phosphatase 2B (PP2B, calcineurin) in Paramecium: partial characterization reveals that two members of the unusually large catalytic subunit family have distinct roles in calcium-dependent processes.Eukaryot Cell. 2010 Jul;9(7):1049-63. doi: 10.1128/EC.00322-09. Epub 2010 Apr 30. Eukaryot Cell. 2010. PMID: 20435698 Free PMC article.
-
A compendium of human gene functions derived from evolutionary modelling.Nature. 2025 Apr;640(8057):146-154. doi: 10.1038/s41586-025-08592-0. Epub 2025 Feb 26. Nature. 2025. PMID: 40011791 Free PMC article.
-
Role of Ectopic Gene Conversion in the Evolution of a Candida krusei Pleiotropic Drug Resistance Transporter Family.Genetics. 2017 Apr;205(4):1619-1639. doi: 10.1534/genetics.116.194811. Epub 2017 Feb 3. Genetics. 2017. PMID: 28159755 Free PMC article.
-
Genome-wide identification and analysis of late embryogenesis abundant (LEA) genes in Prunus mume.Mol Biol Rep. 2013 Feb;40(2):1937-46. doi: 10.1007/s11033-012-2250-3. Epub 2012 Oct 21. Mol Biol Rep. 2013. PMID: 23086279
-
Gene tree species tree reconciliation with gene conversion.J Math Biol. 2019 May;78(6):1981-2014. doi: 10.1007/s00285-019-01331-w. Epub 2019 Feb 15. J Math Biol. 2019. PMID: 30767052
References
-
- Ohno S. Evolution by Gene Duplication. New York: Springer-Verlag; 1970.
-
- Petes TD, Hill CW. Recombination between repeated genes in microorganisms. Annu Rev Genet. 1988;22:147–168. - PubMed
-
- Ohta T. Evolution and Variation of Multigene Families. Berlin/New York: Springer-Verlag; 1980.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

