Haploinsufficiency of activation-induced deaminase for antibody diversification and chromosome translocations both in vitro and in vivo
- PMID: 19079594
- PMCID: PMC2592691
- DOI: 10.1371/journal.pone.0003927
Haploinsufficiency of activation-induced deaminase for antibody diversification and chromosome translocations both in vitro and in vivo
Abstract
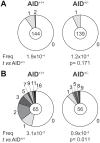
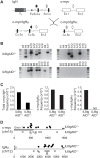
The humoral immune response critically relies on the secondary diversification of antibodies. This diversification takes places through somatic remodelling of the antibody genes by two molecular mechanisms, Class Switch Recombination (CSR) and Somatic Hypermutation (SHM). The enzyme Activation Induced Cytidine Deaminase (AID) initiates both SHM and CSR by deaminating cytosine residues on the DNA of immunoglobulin genes. While crucial for immunity, AID-catalysed deamination is also the triggering event for the generation of lymphomagenic chromosome translocations. To address whether restricting the levels of AID expression in vivo contributes to the regulation of its function, we analysed mice harbouring a single copy of the AID gene (AID(+/-)). AID(+/-) mice express roughly 50% of normal AID levels, and display a mild hyperplasia, reminiscent of AID deficient mice and humans. Moreover, we found that AID(+/-) cells have an impaired competence for CSR and SHM, which indicates that AID gene dose is limiting for its physiologic function. We next evaluated the impact of AID reduction in AID(+/-) mice on the generation of chromosome translocations. Our results show that the frequency of AID-promoted c-myc/IgH translocations is reduced in AID(+/-) mice, both in vivo and in vitro. Therefore, AID is haploinsufficient for antibody diversification and chromosome translocations. These findings suggest that limiting the physiologic levels of AID expression can be a regulatory mechanism that ensures an optimal balance between immune proficiency and genome integrity.
Conflict of interest statement
Figures


References
-
- Schatz DG. V(D)J recombination. Immunol Rev. 2004;200:5–11. - PubMed
-
- Peled JU, Kuang FL, Iglesias-Ussel MD, Roa S, Kalis SL, et al. The biochemistry of somatic hypermutation. Annu Rev Immunol. 2008;26:481–511. - PubMed
-
- Di Noia JM, Neuberger MS. Molecular mechanisms of antibody somatic hypermutation. Annu Rev Biochem. 2007;76:1–22. - PubMed
-
- Stavnezer J. Antibody class switching. Adv Immunol. 1996;61:79–146. - PubMed
-
- Stavnezer J. Molecular processes that regulate class switching. Curr Top Microbiol Immunol. 2000;245:127–168. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

