Bile acids activate fibroblast growth factor 19 signaling in human hepatocytes to inhibit cholesterol 7alpha-hydroxylase gene expression
- PMID: 19085950
- PMCID: PMC2614454
- DOI: 10.1002/hep.22627
Bile acids activate fibroblast growth factor 19 signaling in human hepatocytes to inhibit cholesterol 7alpha-hydroxylase gene expression
Abstract
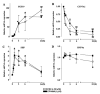
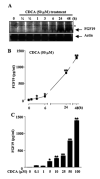
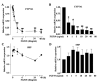
Mouse fibroblast growth factor 15 (FGF15) and human ortholog FGF19 have been identified as the bile acid-induced intestinal factors that mediate bile acid feedback inhibition of cholesterol 7alpha-hydroxylase gene (C YP7A1) transcription in mouse liver. The mechanism underlying FGF15/FGF19 inhibition of bile acid synthesis in hepatocytes remains unclear. Chenodeoxycholic acid (CDCA) and the farnesoid X receptor (FXR)-specific agonist GW4064 strongly induced FGF19 but inhibited CYP7A1 messenger RNA (mRNA) levels in primary human hepatocytes. FGF19 strongly and rapidly repressed CYP7A1 but not small heterodimer partner (SHP) mRNA levels. Kinase inhibition and phosphorylation assays revealed that the mitogen-activated protein kinase/extracellular signal-regulated kinase 1/2 (MAPK/Erk1/2) pathway played a major role in mediating FGF19 inhibition of CYP7A1. However, small interfering RNA (siRNA) knockdown of SHP did not affect FGF19 inhibition of CYP7A1. Interestingly, CDCA stimulated tyrosine phosphorylation of the FGF receptor 4 (FGFR4) in hepatocytes. FGF19 antibody and siRNA specific to FGFR4 abrogated GW4064 inhibition of CYP7A1. These results suggest that bile acid-activated FXR is able to induce FGF19 in hepatocytes to inhibit CYP7A1 by an autocrine/paracrine mechanism.
Conclusion: The hepatic FGF19/FGFR4/Erk1/2 pathway may inhibit CYP7A1 independent of SHP. In addition to inducing FGF19 in the intestine, bile acids in hepatocytes may activate the liver FGF19/FGFR4 signaling pathway to inhibit bile acid synthesis and prevent accumulation of toxic bile acid in human livers.
Conflict of interest statement
Potential conflict of interest: nothing to report.
Figures
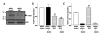
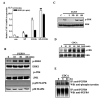

References
-
- Myant NB, Mitropoulos KA. Cholesterol 7 alpha-hydroxylase. J Lipid Res. 1977;18:135–153. - PubMed
-
- Eriksson S. Biliary excretion of bile acids and cholesterol in bile fistula rats; bile acids and steroids. Proc Soc Exp Biol Med. 1957;94:578–582. - PubMed
-
- Bergstrom S, Danielsson H. On the regulation of bile acid formation in the rat liver. Acta Physiol Scand. 1958;43:1–7. - PubMed
-
- Shefer S, Hauser S, Bekersky I, Mosbach EH. Biochemical site of regulation of bile acid biosynthesis in the rat. J Lipid Res. 1970;11:404–411. - PubMed
-
- Russell DW. The enzymes, regulation, and genetics of bile acid synthesis. Annu Rev Biochem. 2003;72:137–174. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous
