Flanking signal and mature peptide residues influence signal peptide cleavage
- PMID: 19091014
- PMCID: PMC2638155
- DOI: 10.1186/1471-2105-9-S12-S15
Flanking signal and mature peptide residues influence signal peptide cleavage
Abstract
Background: Signal peptides (SPs) mediate the targeting of secretory precursor proteins to the correct subcellular compartments in prokaryotes and eukaryotes. Identifying these transient peptides is crucial to the medical, food and beverage and biotechnology industries yet our understanding of these peptides remains limited. This paper examines the most common type of signal peptides cleavable by the endoprotease signal peptidase I (SPase I), and the residues flanking the cleavage sites of three groups of signal peptide sequences, namely (i) eukaryotes (Euk) (ii) Gram-positive (Gram+) bacteria, and (iii) Gram-negative (Gram-) bacteria.
Results: In this study, 2352 secretory peptide sequences from a variety of organisms with amino-terminal SPs are extracted from the manually curated SPdb database for analysis based on physicochemical properties such as pI, aliphatic index, GRAVY score, hydrophobicity, net charge and position-specific residue preferences. Our findings show that the three groups share several similarities in general, but they display distinctive features upon examination in terms of their amino acid compositions and frequencies, and various physico-chemical properties. Thus, analysis or prediction of their sequences should be separated and treated as distinct groups.
Conclusion: We conclude that the peptide segment recognized by SPase I extends to the start of the mature protein to a limited extent, upon our survey of the amino acid residues surrounding the cleavage processing site. These flanking residues possibly influence the cleavage processing and contribute to non-canonical cleavage sites. Our findings are applicable in defining more accurate prediction tools for recognition and identification of cleavage site of SPs.
Figures

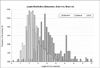
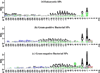

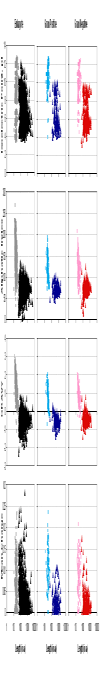
References
-
- Pugsley AP. Protein targeting. San Diego; London: Academic Press; 1989.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

