Using a kernel density estimation based classifier to predict species-specific microRNA precursors
- PMID: 19091019
- PMCID: PMC2638167
- DOI: 10.1186/1471-2105-9-S12-S2
Using a kernel density estimation based classifier to predict species-specific microRNA precursors
Abstract
Background: MicroRNAs (miRNAs) are short non-coding RNA molecules participating in post-transcriptional regulation of gene expression. There have been many efforts to discover miRNA precursors (pre-miRNAs) over the years. Recently, ab initio approaches obtain more attention because that they can discover species-specific pre-miRNAs. Most ab initio approaches proposed novel features to characterize RNA molecules. However, there were fewer discussions on the associated classification mechanism in a miRNA predictor.
Results: This study focuses on the classification algorithm for miRNA prediction. We develop a novel ab initio method, miR-KDE, in which most of the features are collected from previous works. The classification mechanism in miR-KDE is the relaxed variable kernel density estimator (RVKDE) that we have recently proposed. When compared to the famous support vector machine (SVM), RVKDE exploits more local information of the training dataset. MiR-KDE is evaluated using a training set consisted of only human pre-miRNAs to predict a benchmark collected from 40 species. The experimental results show that miR-KDE delivers favorable performance in predicting human pre-miRNAs and has advantages for pre-miRNAs from the genera taxonomically distant to humans.
Conclusion: We use a novel classifier of which the characteristic of exploiting local information is particularly suitable to predict species-specific pre-miRNAs. This study also provides a comprehensive analysis from the view of classification mechanism. The good performance of miR-KDE encourages more efforts on the classification methodology as well as the feature extraction in miRNA prediction.
Figures
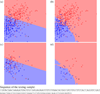
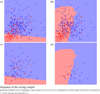
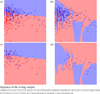
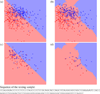

Similar articles
-
Predicting microRNA precursors with a generalized Gaussian components based density estimation algorithm.BMC Bioinformatics. 2010 Jan 18;11 Suppl 1(Suppl 1):S52. doi: 10.1186/1471-2105-11-S1-S52. BMC Bioinformatics. 2010. PMID: 20122227 Free PMC article.
-
Classification of real and pseudo microRNA precursors using local structure-sequence features and support vector machine.BMC Bioinformatics. 2005 Dec 29;6:310. doi: 10.1186/1471-2105-6-310. BMC Bioinformatics. 2005. PMID: 16381612 Free PMC article.
-
Predicting human microRNA precursors based on an optimized feature subset generated by GA-SVM.Genomics. 2011 Aug;98(2):73-8. doi: 10.1016/j.ygeno.2011.04.011. Epub 2011 May 14. Genomics. 2011. PMID: 21586321
-
Popular Computational Tools Used for miRNA Prediction and Their Future Development Prospects.Interdiscip Sci. 2020 Dec;12(4):395-413. doi: 10.1007/s12539-020-00387-3. Epub 2020 Sep 21. Interdiscip Sci. 2020. PMID: 32959233 Review.
-
Predicting novel microRNA: a comprehensive comparison of machine learning approaches.Brief Bioinform. 2019 Sep 27;20(5):1607-1620. doi: 10.1093/bib/bby037. Brief Bioinform. 2019. PMID: 29800232 Review.
Cited by
-
Genome-wide approaches in the study of microRNA biology.Wiley Interdiscip Rev Syst Biol Med. 2011 Sep-Oct;3(5):491-512. doi: 10.1002/wsbm.128. Epub 2010 Dec 31. Wiley Interdiscip Rev Syst Biol Med. 2011. PMID: 21197653 Free PMC article. Review.
-
A fast ab-initio method for predicting miRNA precursors in genomes.Nucleic Acids Res. 2012 Jun;40(11):e80. doi: 10.1093/nar/gks146. Epub 2012 Feb 22. Nucleic Acids Res. 2012. PMID: 22362754 Free PMC article.
-
ViralmiR: a support-vector-machine-based method for predicting viral microRNA precursors.BMC Bioinformatics. 2015;16 Suppl 1(Suppl 1):S9. doi: 10.1186/1471-2105-16-S1-S9. Epub 2015 Jan 21. BMC Bioinformatics. 2015. PMID: 25708359 Free PMC article.
-
Predicting microRNA precursors with a generalized Gaussian components based density estimation algorithm.BMC Bioinformatics. 2010 Jan 18;11 Suppl 1(Suppl 1):S52. doi: 10.1186/1471-2105-11-S1-S52. BMC Bioinformatics. 2010. PMID: 20122227 Free PMC article.
-
Emerging strengths in Asia Pacific bioinformatics.BMC Bioinformatics. 2008 Dec 12;9 Suppl 12(Suppl 12):S1. doi: 10.1186/1471-2105-9-S12-S1. BMC Bioinformatics. 2008. PMID: 19091008 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

