Gene expression levels are a target of recent natural selection in the human genome
- PMID: 19091723
- PMCID: PMC2767089
- DOI: 10.1093/molbev/msn289
Gene expression levels are a target of recent natural selection in the human genome
Abstract
Changes in gene expression may represent an important mode of human adaptation. However, to date, there are relatively few known examples in which selection has been shown to act directly on levels or patterns of gene expression. In order to test whether single nucleotide polymorphisms (SNPs) that affect gene expression in cis are frequently targets of positive natural selection in humans, we analyzed genome-wide SNP and expression data from cell lines associated with the International HapMap Project. Using a haplotype-based test for selection that was designed to detect incomplete selective sweeps, we found that SNPs showing signals of selection are more likely than random SNPs to be associated with gene expression levels in cis. This signal is significant in the Yoruba (which is the population that shows the strongest signals of selection overall) and shows a trend in the same direction in the other HapMap populations. Our results argue that selection on gene expression levels is an important type of human adaptation. Finally, our work provides an analytical framework for tackling a more general problem that will become increasingly important: namely, testing whether selection signals overlap significantly with SNPs that are associated with phenotypes of interest.
Figures
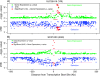
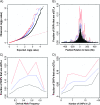
References
-
- Altshuler D, Hirschhorn J, Klannemark M, et al. (16 co-authors) The common PPARgamma Pro12Ala polymorphism is associated with decreased risk of type 2 diabetes. Nat Genet. 2000;26:76–80. - PubMed
-
- Dixon A, Liang L, Moffatt M, et al. (13 co-authors) A genome-wide association study of global gene expression. Nat Genet. 2007;39:1202–1207. - PubMed
-
- Enattah N, Forsblom C, Rasinper H, Tuomi T, Groop P, Jrvel I. The genetic variant of lactase persistence C (-13910) T as a risk factor for type I and II diabetes in the Finnish population. Eur J Clin Nutr. 2004;58:1319–1322. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

