Telomere elongation involves intra-molecular DNA replication in cells utilizing alternative lengthening of telomeres
- PMID: 19095716
- PMCID: PMC2649016
- DOI: 10.1093/hmg/ddn436
Telomere elongation involves intra-molecular DNA replication in cells utilizing alternative lengthening of telomeres
Abstract
Alternative lengthening of telomeres (ALT) is a telomere length maintenance mechanism based on recombination, where telomeres use other telomeric DNA as a template for DNA synthesis. About 10% of all human tumors depend on ALT for their continued growth, and understanding its molecular details is critically important for the development of cancer treatments that target this mechanism. We have previously shown that telomeres of ALT-positive human cells can become lengthened via inter-telomeric copying, i.e. by copying the telomere of another chromosome. The possibility that such telomeres could elongate by using other sources of telomeric DNA as copy templates has not been investigated previously. In this study, we have determined whether a telomere can become lengthened by copying its own sequences, without the need for using another telomere as a copy template. To test this, we transduced an ALT cell line with a telomere-targeting construct and obtained clones with a single tagged telomere. We showed that the telomere tag can be amplified without the involvement of other telomeres, indicating that telomere elongation can also occur by intra-telomeric DNA copying. This is the first direct evidence that the ALT mechanism involves more than one method of telomere elongation.
Figures


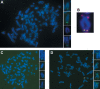




References
-
- Yeager T.R., Neumann A.A., Englezou A., Huschtscha L.I., Noble J.R., Reddel R.R. Telomerase-negative immortalized human cells contain a novel type of promyelocytic leukemia (PML) body. Cancer Res. 1999;59:4175–4179. - PubMed
-
- Dunham M.A., Neumann A.A., Fasching C.L., Reddel R.R. Telomere maintenance by recombination in human cells. Nat. Genet. 2000;26:447–450. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

