Conservation of mechanisms mediating gonadotrophin-releasing hormone 1 stimulation of human luteinizing hormone beta subunit transcription
- PMID: 19106114
- PMCID: PMC2734162
- DOI: 10.1093/molehr/gan079
Conservation of mechanisms mediating gonadotrophin-releasing hormone 1 stimulation of human luteinizing hormone beta subunit transcription
Abstract
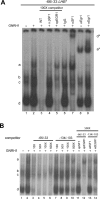
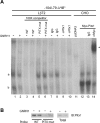
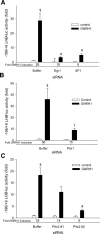
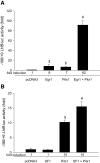
Gonadotrophin-releasing hormone (GNRH1) regulates pituitary luteinizing hormone (LH). Previous studies have delineated a mechanism for GNRH1-induced LHbeta subunit gene (Lhb) transcription, the rate-limiting step in LH production. GNRH1 induces expression of early growth response 1 (EGR1), which interacts with steroidogenic factor 1 (SF1) and paired-like homeodomain transcription factor 1 (PITX1) to regulate Lhb promoter activity. Though the cis-elements for these factors are conserved across species, regulation of human LHB transcription has not been thoroughly investigated. We therefore characterized LHB transcriptional regulation by GNRH1 using promoter-reporter analyses in LbetaT2 cells. GNRH1 stimulated LHB transcription via an extracellular signal-regulated kinase 1/2 pathway. EGR1 bound to two binding sites on the LHB promoter and this binding was increased by GNRH1. Mutation of either site or knockdown of endogenous EGR1 decreased basal and/or GNRH1-regulated promoter activity. The human LHB promoter also contains low and high affinity SF1 binding sites. Mutation of these elements or depletion of endogenous SF1 impaired basal and ligand-induced transcription. Knockdown of PITX1 or PITX2 isoforms impaired GNRH1 induction, and endogenous PITX1 bound to the candidate PITX binding site on the LHB promoter. Thus, the mechanism described for GNRH1 regulation of Lhb in other species is largely conserved for human LHB. We also uncover a previously unappreciated role for PITX2 isoforms in this process.
Figures


Similar articles
-
SMAD3 and EGR1 physically and functionally interact in promoter-specific fashion.Cell Signal. 2010 Jun;22(6):936-43. doi: 10.1016/j.cellsig.2010.01.019. Epub 2010 Feb 8. Cell Signal. 2010. PMID: 20149866
-
Maximal activity of the luteinizing hormone beta-subunit gene requires beta-catenin.Mol Endocrinol. 2007 Apr;21(4):963-71. doi: 10.1210/me.2006-0383. Epub 2007 Jan 23. Mol Endocrinol. 2007. PMID: 17244763
-
Luteinizing hormone beta promoter stimulation by adenylyl cyclase and cooperation with gonadotropin-releasing hormone 1 in transgenic mice and LBetaT2 Cells.Biol Reprod. 2007 Dec;77(6):1073-80. doi: 10.1095/biolreprod.107.064139. Epub 2007 Aug 15. Biol Reprod. 2007. PMID: 17699734
-
Mechanisms for pulsatile regulation of the gonadotropin subunit genes by GNRH1.Biol Reprod. 2006 Jun;74(6):993-8. doi: 10.1095/biolreprod.105.049049. Epub 2006 Feb 15. Biol Reprod. 2006. PMID: 16481592 Review.
-
Transcriptional regulation of the LH beta gene by gonadotropin-releasing hormone and the protein kinase C system.Vitam Horm. 2000;60:195-227. doi: 10.1016/s0083-6729(00)60020-1. Vitam Horm. 2000. PMID: 11037625 Review. No abstract available.
Cited by
-
Gonadotropin-releasing hormone regulates transcription of the inhibin B co-receptor, TGFBR3L, via early growth response one.J Biol Chem. 2025 Apr;301(4):108405. doi: 10.1016/j.jbc.2025.108405. Epub 2025 Mar 14. J Biol Chem. 2025. PMID: 40090584 Free PMC article.
-
Genetic regulation of murine pituitary development.J Mol Endocrinol. 2015 Apr;54(2):R55-73. doi: 10.1530/JME-14-0237. Epub 2015 Jan 13. J Mol Endocrinol. 2015. PMID: 25587054 Free PMC article. Review.
-
GnRH pulsatility, the pituitary response and reproductive dysfunction.Endocr J. 2009;56(6):729-37. doi: 10.1507/endocrj.k09e-185. Epub 2009 Jul 17. Endocr J. 2009. PMID: 19609045 Free PMC article. Review.
-
Murine FSH Production Depends on the Activin Type II Receptors ACVR2A and ACVR2B.Endocrinology. 2020 Jul 1;161(7):bqaa056. doi: 10.1210/endocr/bqaa056. Endocrinology. 2020. PMID: 32270195 Free PMC article.
-
CREB binding protein (CBP) activation is required for luteinizing hormone beta expression and normal fertility in mice.Mol Cell Biol. 2012 Jul;32(13):2349-58. doi: 10.1128/MCB.00394-12. Epub 2012 Apr 16. Mol Cell Biol. 2012. PMID: 22508984 Free PMC article.
References
-
- Bernard DJ. Both SMAD2 and SMAD3 mediate activin-stimulated expression of the follicle-stimulating hormone beta subunit in mouse gonadotrope cells. Mol Endocrinol. 2004;18:606–623. - PubMed
-
- Call GB, Wolfe MW. Species differences in GnRH activation of the LHbeta promoter: role of Egr1 and Sp1. Mol Cell Endocrinol. 2002;189:85–96. - PubMed
-
- Chaney BA, Clark-Baldwin K, Dave V, Ma J, Rance M. Solution structure of the K50 class homeodomain PITX2 bound to DNA and implications for mutations that cause Rieger syndrome. Biochemistry. 2005;44:7497–7511. - PubMed
-
- Dorn C, Ou Q, Svaren J, Crawford PA, Sadovsky Y. Activation of luteinizing hormone beta gene by gonadotropin-releasing hormone requires the synergy of early growth response-1 and steroidogenic factor-1. J Biol Chem. 1999;274:13870–13876. - PubMed

