Fine mapping on chromosome 10q22-q23 implicates Neuregulin 3 in schizophrenia
- PMID: 19118813
- PMCID: PMC2668048
- DOI: 10.1016/j.ajhg.2008.12.005
Fine mapping on chromosome 10q22-q23 implicates Neuregulin 3 in schizophrenia
Abstract
Linkage studies have implicated 10q22-q23 as a schizophrenia (SZ) susceptibility locus in Ashkenazi Jewish (AJ) and Han Chinese from Taiwan populations. To further explore our previous linkage signal in the AJ population (NPL score: 4.27, empirical p = 2 x 10(-5)), we performed a peakwide association fine mapping study by using 1414 SNPs across approximately 12.5 Mb in 10q22-q23. We genotyped 1515 AJ individuals, including 285 parent-child trios, 173 unrelated cases, and 487 unrelated controls. We analyzed the binary diagnostic phenotype of SZ and 9 heritable quantitative traits derived from a principal components factor analysis of 73 items from our consensus diagnostic ratings and direct assessment interviews. Although no marker withstood multiple test correction for association with the binary SZ phenotype, we found strong evidence of association by using the "delusion" factor as the quantitative trait at three SNPs (rs10883866, rs10748842, and rs6584400) located in a 13 kb interval in intron 1 of Neuregulin 3 (NRG3). Our best p value from family-based association analysis was 7.26 x 10(-7). We replicated this association in the collection of 173 unrelated AJ cases (p = 1.55 x 10(-2)), with a combined p value of 2.30 x 10(-7). After performing 10,000 permutations of each of the phenotypes, we estimated the empirical study-wide significance across all 9 factors (90,000 permutations) to be p = 2.7 x 10(-3). NRG3 is primarily expressed in the central nervous system and is one of three paralogs of NRG1, a gene strongly implicated in SZ. These biological properties together with our linkage and association results strongly support NRG3 as a gene involved in SZ.
Figures

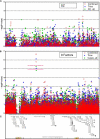


References
-
- Eaton W.W. Epidemiology of schizophrenia. Epidemiol. Rev. 1985;7:105–126. - PubMed
-
- McGue M., Gottesman I.I. The genetic epidemiology of schizophrenia and the design of linkage studies. Eur. Arch. Psychiatry Clin. Neurosci. 1991;240:174–181. - PubMed
-
- Tsuang M.T., Gilbertson M.W., Faraone S.V. The genetics of schizophrenia. Current knowledge and future directions. Schizophr. Res. 1991;4:157–171. - PubMed
-
- Cardno A.G., Gottesman I.I. Twin studies of schizophrenia: from bow-and-arrow concordances to star wars Mx and functional genomics. Am. J. Med. Genet. 2000;97:12–17. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

