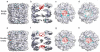
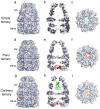
Chaperonin complex with a newly folded protein encapsulated in the folding chamber
- PMID: 19122642
- PMCID: PMC2728927
- DOI: 10.1038/nature07479
Chaperonin complex with a newly folded protein encapsulated in the folding chamber
Abstract
A subset of essential cellular proteins requires the assistance of chaperonins (in Escherichia coli, GroEL and GroES), double-ring complexes in which the two rings act alternately to bind, encapsulate and fold a wide range of nascent or stress-denatured proteins. This process starts by the trapping of a substrate protein on hydrophobic surfaces in the central cavity of a GroEL ring. Then, binding of ATP and co-chaperonin GroES to that ring ejects the non-native protein from its binding sites, through forced unfolding or other major conformational changes, and encloses it in a hydrophilic chamber for folding. ATP hydrolysis and subsequent ATP binding to the opposite ring trigger dissociation of the chamber and release of the substrate protein. The bacteriophage T4 requires its own version of GroES, gp31, which forms a taller folding chamber, to fold the major viral capsid protein gp23 (refs 16-20). Polypeptides are known to fold inside the chaperonin complex, but the conformation of an encapsulated protein has not previously been visualized. Here we present structures of gp23-chaperonin complexes, showing both the initial captured state and the final, close-to-native state with gp23 encapsulated in the folding chamber. Although the chamber is expanded, it is still barely large enough to contain the elongated gp23 monomer, explaining why the GroEL-GroES complex is not able to fold gp23 and showing how the chaperonin structure distorts to enclose a large, physiological substrate protein.
Figures


References
-
- Sigler PB, et al. Structure and function in GroEL-mediated protein folding. Annu. Rev. Biochem. 1998;67:581. - PubMed
-
- Rye HS, et al. GroEL-GroES cycling: ATP and nonnative polypeptide direct alternation of folding-active rings. Cell. 1999;97:325. - PubMed
-
- Houry WA, et al. Identification of in vivo substrates of the chaperonin GroEL. Nature. 1999;402:147. - PubMed
-
- Kerner MJ, et al. Proteome-wide analysis of chaperonin-dependent protein folding in Escherichia coli. Cell. 2005;122:209. - PubMed
-
- Horwich AL, Fenton WA, Chapman E, Farr GW. Two families of chaperonin: physiology and mechanism. Annu Rev Cell Dev Biol. 2007;23:115. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

