Transcription is required for establishment of germline methylation marks at imprinted genes
- PMID: 19136628
- PMCID: PMC2632167
- DOI: 10.1101/gad.495809
Transcription is required for establishment of germline methylation marks at imprinted genes
Erratum in
- Genes Dev. 2009 Oct 1;23(19):2358
Abstract
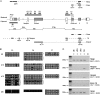
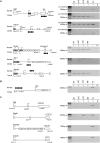
Genomic imprinting requires the differential marking by DNA methylation of genes in male and female gametes. In the female germline, acquisition of methylation imprint marks depends upon the de novo methyltransferase Dnmt3a and its cofactor Dnmt3L, but the reasons why specific sequences are targets for Dnmt3a and Dnmt3L are still poorly understood. Here, we investigate the role of transcription in establishing maternal germline methylation marks. We show that at the Gnas locus, truncating transcripts from the furthest upstream Nesp promoter disrupts oocyte-derived methylation of the differentially methylated regions (DMRs). Transcription through DMRs in oocytes is not restricted to this locus but occurs across the prospective DMRs at many other maternally marked imprinted domains, suggesting a common requirement for transcription events. The transcripts implicated here in gametic methylation are protein-coding, in contrast to the noncoding antisense transcripts involved in the monoallelic silencing of imprinted genes in somatic tissues, although they often initiate from alternative promoters in oocytes. We propose that transcription is a third essential component of the de novo methylation system, which includes optimal CpG spacing and histone modifications, and may be required to create or maintain open chromatin domains to allow the methylation complex access to its preferred targets.
Figures



References
-
- Arnaud P., Monk D., Hitchins M., Gordon E., Dean W., Beechey C.V., Peters J., Craigen W., Preece M., Stanier P., et al. Conserved methylation imprints in the human and mouse GRB10 genes with divergent allelic expression suggests differential reading of the same mark. Hum. Mol. Genet. 2003;12:1005–1019. - PubMed
-
- Arnaud P., Hata K., Kaneda M., Li E., Sasaki H., Feil R., Kelsey G. Stochastic imprinting in the progeny of Dnmt3L−/− females. Hum. Mol. Genet. 2006;15:589–598. - PubMed
-
- Bastepe M., Frohlich L.F., Hendy G.N., Indridason O.S., Josse R.G., Koshiyama H., Korkko J., Nakamoto J.M., Rosenbloom A.L., Slyper A.H., et al. Autosomal dominant pseudohypoparathyroidism type Ib is associated with a heterozygous microdeletion that likely disrupts a putative imprinting control element of GNAS. J. Clin. Invest. 2003;112:1255–1263. - PMC - PubMed
-
- Bastepe M., Frohlich L.F., Linglart A., Abu-Zahra H.S., Tojo K., Ward L.M., Juppner H. Deletion of the NESP55 differentially methylated region causes loss of maternal GNAS imprints and pseudohypoparathyroidism type Ib. Nat. Genet. 2005;37:25–27. - PubMed
-
- Bernstein B.E., Kamal M., Lindblad-Toh K., Bekiranov S., Bailey D.K., Huebert D.J., McMahon S., Karlsson E.K., Kulbokas E.J., III, Gingeras T.R., et al. Genomic maps and comparative analysis of histone modifications in human and mouse. Cell. 2005;120:169–181. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
