An expanded clade of rodent Trim5 genes
- PMID: 19147168
- PMCID: PMC2692226
- DOI: 10.1016/j.virol.2008.12.018
An expanded clade of rodent Trim5 genes
Abstract
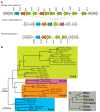
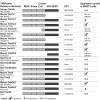
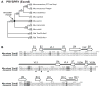
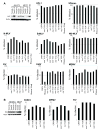
Trim5alpha from primates (including humans), cows, and rabbits has been shown to be an active antiviral host gene that acts against a range of retroviruses. Although this suggests that Trim5alpha may be a common antiviral restriction factor among mammals, the status of Trim5 genes in rodents has been unclear. Using genomic and phylogenetic analyses, we describe an expanded paralogous cluster of at least eight Trim5-like genes in mice (including the previously described Trim12 and Trim30 genes), and three Trim5-like genes in rats. Our characterization of the rodent Trim5 locus, and comparison to the Trim5 locus in humans, cows, and rabbits, indicates that Trim5 has undergone independent evolutionary expansions within species. Evolutionary analysis shows that rodent Trim5 genes have evolved under positive selection, suggesting evolutionary conflicts consistent with important antiviral function. Sampling six rodent Trim5 genes failed to reveal antiviral activities against a set of eight retroviral challenges, although we predict that such activities exist.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures


Similar articles
-
The B30.2(SPRY) domain of the retroviral restriction factor TRIM5alpha exhibits lineage-specific length and sequence variation in primates.J Virol. 2005 May;79(10):6111-21. doi: 10.1128/JVI.79.10.6111-6121.2005. J Virol. 2005. PMID: 15857996 Free PMC article.
-
Discordant evolution of the adjacent antiretroviral genes TRIM22 and TRIM5 in mammals.PLoS Pathog. 2007 Dec;3(12):e197. doi: 10.1371/journal.ppat.0030197. PLoS Pathog. 2007. PMID: 18159944 Free PMC article.
-
Truncation of TRIM5 in the Feliformia explains the absence of retroviral restriction in cells of the domestic cat.J Virol. 2009 Aug;83(16):8270-5. doi: 10.1128/JVI.00670-09. Epub 2009 Jun 3. J Virol. 2009. PMID: 19494015 Free PMC article.
-
The control of viral infection by tripartite motif proteins and cyclophilin A.Retrovirology. 2007 Jun 12;4:40. doi: 10.1186/1742-4690-4-40. Retrovirology. 2007. PMID: 17565686 Free PMC article. Review.
-
The cell biology of TRIM5α.Curr HIV/AIDS Rep. 2012 Mar;9(1):73-80. doi: 10.1007/s11904-011-0102-8. Curr HIV/AIDS Rep. 2012. PMID: 22193888 Free PMC article. Review.
Cited by
-
Evolution-guided functional analyses reveal diverse antiviral specificities encoded by IFIT1 genes in mammals.Elife. 2016 May 31;5:e14228. doi: 10.7554/eLife.14228. Elife. 2016. PMID: 27240734 Free PMC article.
-
Long-term balancing selection maintains trans-specific polymorphisms in the human TRIM5 gene.Hum Genet. 2010 Dec;128(6):577-88. doi: 10.1007/s00439-010-0884-6. Epub 2010 Sep 2. Hum Genet. 2010. PMID: 20811909
-
Highly-potent, synthetic APOBEC3s restrict HIV-1 through deamination-independent mechanisms.PLoS Pathog. 2021 Jun 25;17(6):e1009523. doi: 10.1371/journal.ppat.1009523. eCollection 2021 Jun. PLoS Pathog. 2021. PMID: 34170969 Free PMC article.
-
Elucidation of the Complicated Scenario of Primate APOBEC3 Gene Evolution.J Virol. 2021 May 24;95(12):e00144-21. doi: 10.1128/JVI.00144-21. Print 2021 May 24. J Virol. 2021. PMID: 33789992 Free PMC article.
-
TRIMmunity: the roles of the TRIM E3-ubiquitin ligase family in innate antiviral immunity.J Mol Biol. 2014 Mar 20;426(6):1265-84. doi: 10.1016/j.jmb.2013.12.005. Epub 2013 Dec 12. J Mol Biol. 2014. PMID: 24333484 Free PMC article. Review.
References
-
- Bartz SR, Vodicka MA. Production of high-titer human immunodeficiency virus type 1 pseudotyped with vesicular stomatitis virus glycoprotein. Methods. 1997;12:337–342. - PubMed
-
- Best S, Le Tissier P, Towers G, Stoye JP. Positional cloning of the mouse retrovirus restriction gene Fv1. Nature. 1996;382:826–829. - PubMed
-
- Bieniasz PD. Intrinsic immunity: A front-line defense against viral attack. Nat immunol. 2004;5:1109–1115. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

