Protein arginine methylation in mammals: who, what, and why
- PMID: 19150423
- PMCID: PMC3372459
- DOI: 10.1016/j.molcel.2008.12.013
Protein arginine methylation in mammals: who, what, and why
Abstract
The covalent marking of proteins by methyl group addition to arginine residues can promote their recognition by binding partners or can modulate their biological activity. A small family of gene products that catalyze such methylation reactions in eukaryotes (PRMTs) works in conjunction with a changing cast of associated subunits to recognize distinct cellular substrates. These reactions display many of the attributes of reversible covalent modifications such as protein phosphorylation or protein lysine methylation; however, it is unclear to what extent protein arginine demethylation occurs. Physiological roles for protein arginine methylation have been established in signal transduction, mRNA splicing, transcriptional control, DNA repair, and protein translocation.
Figures
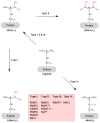

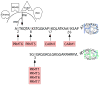
References
-
- Adams MM, Wang B, Xia Z, Morales JC, Lu X, Donehower LA, Bochar DA, Elledge SJ, Carpenter PB. 53BP1 oligomerization is independent of its methylation by PRMT1. Cell Cycle. 2005;4:1854–1861. - PubMed
-
- Ancelin K, Lange UC, Hajkova P, Schneider R, Bannister AJ, Kouzarides T, Surani MA. Blimp1 associates with Prmt5 and directs histone arginine methylation in mouse germ cells. Nat Cell Biol. 2006;8:623–630. - PubMed
-
- Bedford MT. Arginine methylation at a glance. J Cell Sci. 2007;120:4243–4246. - PubMed
-
- Bedford MT, Frankel A, Yaffe MB, Clarke S, Leder P, Richard S. Arginine methylation inhibits the binding of proline-rich ligands to Src homology 3, but not WW, domains. J Biol Chem. 2000;275:16030–16036. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

