GRIM-19 inhibits v-Src-induced cell motility by interfering with cytoskeletal restructuring
- PMID: 19151760
- PMCID: PMC2662763
- DOI: 10.1038/onc.2008.480
GRIM-19 inhibits v-Src-induced cell motility by interfering with cytoskeletal restructuring
Abstract
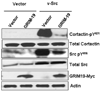
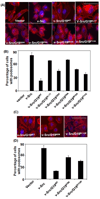
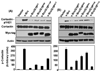
GRIM-19 (Gene associated with Retinoid-Interferon-induced Mortality 19) is a novel tumor suppressor regulated by interferon/retinoid combination. We have recently shown that GRIM-19 inhibits v-Src-induced oncogenic transformation and metastatic behavior of cells. Oncogenic v-Src induces cell motility by cytoskeletal remodeling, especially the formation of podosomes and. Here, we show that GRIM-19 inhibited the v-Src-induced cell motility by inhibiting cytoskeletal remodeling, that is, podosome formation. We also show that the N terminus of GRIM-19 played a major role in this process and identified critical residues in this region. More importantly, we show that tumor-associated GRIM-19 mutations disrupted its ability to inhibit v-Src-induced cell motility. These actions appear to occur independently of STAT3, a known target of GRIM-19-mediated inhibition. Lastly, tumor-associated GRIM-19 mutants significantly lost their ability to control v-Src-induced metastases in vivo, indicating the biological and pathological significance of these observations.
Figures


Similar articles
-
GRIM-19 mutations fail to inhibit v-Src-induced oncogenesis.Oncogene. 2014 Jun 12;33(24):3195-204. doi: 10.1038/onc.2013.271. Epub 2013 Jul 15. Oncogene. 2014. PMID: 23851499 Free PMC article.
-
Tumor suppressive protein gene associated with retinoid-interferon-induced mortality (GRIM)-19 inhibits src-induced oncogenic transformation at multiple levels.Am J Pathol. 2007 Oct;171(4):1352-68. doi: 10.2353/ajpath.2007.070241. Epub 2007 Sep 6. Am J Pathol. 2007. PMID: 17823279 Free PMC article.
-
v-Src rescues actin-based cytoskeletal architecture and cell motility and induces enhanced anchorage independence during oncogenic transformation of focal adhesion kinase-null fibroblasts.J Biol Chem. 2003 Nov 28;278(48):47946-59. doi: 10.1074/jbc.M302720200. Epub 2003 Sep 18. J Biol Chem. 2003. PMID: 14500722
-
GRIM-19: A master regulator of cytokine induced tumor suppression, metastasis and energy metabolism.Cytokine Growth Factor Rev. 2017 Feb;33:1-18. doi: 10.1016/j.cytogfr.2016.09.001. Epub 2016 Sep 15. Cytokine Growth Factor Rev. 2017. PMID: 27659873 Free PMC article. Review.
-
GRIM-19 function in cancer development.Mitochondrion. 2011 Sep;11(5):693-9. doi: 10.1016/j.mito.2011.05.011. Epub 2011 Jun 2. Mitochondrion. 2011. PMID: 21664299 Review.
Cited by
-
GRIM-19 disrupts E6/E6AP complex to rescue p53 and induce apoptosis in cervical cancers.PLoS One. 2011;6(7):e22065. doi: 10.1371/journal.pone.0022065. Epub 2011 Jul 12. PLoS One. 2011. PMID: 21765936 Free PMC article.
-
Glutamine deprivation plus BPTES alters etoposide- and cisplatin-induced apoptosis in triple negative breast cancer cells.Oncotarget. 2016 Aug 23;7(34):54691-54701. doi: 10.18632/oncotarget.10579. Oncotarget. 2016. PMID: 27419628 Free PMC article.
-
Overexpression of GRIM-19, a mitochondrial respiratory chain complex I protein, suppresses hepatocellular carcinoma growth.Int J Clin Exp Pathol. 2014 Oct 15;7(11):7497-507. eCollection 2014. Int J Clin Exp Pathol. 2014. PMID: 25550785 Free PMC article.
-
Cytokine-induced tumor suppressors: a GRIM story.Cytokine. 2010 Oct-Nov;52(1-2):128-42. doi: 10.1016/j.cyto.2010.03.009. Epub 2010 Apr 10. Cytokine. 2010. PMID: 20382543 Free PMC article. Review.
-
Nicotine promotes apoptosis resistance of breast cancer cells and enrichment of side population cells with cancer stem cell-like properties via a signaling cascade involving galectin-3, α9 nicotinic acetylcholine receptor and STAT3.Breast Cancer Res Treat. 2014 May;145(1):5-22. doi: 10.1007/s10549-014-2912-z. Epub 2014 Mar 26. Breast Cancer Res Treat. 2014. PMID: 24668500 Free PMC article.
References
-
- Alchanati I, Nallar SC, Sun P, Gao L, Hu J, Stein A, et al. A proteomic analysis reveals the loss of expression of the cell death regulatory gene GRIM-19 in human renal cell carcinomas. Oncogene. 2006 - PubMed
-
- Angell JE, Lindner DJ, Shapiro PS, Hofmann ER, Kalvakolanu DV. Identification of GRIM-19, a novel cell death-regulatory gene induced by the interferon-beta and retinoic acid combination, using a genetic approach. J Biol Chem. 2000;275:33416–33426. - PubMed
-
- Coussens LM, Fingleton B, Matrisian LM. Matrix metalloproteinase inhibitors and cancer: trials and tribulations. Science. 2002;295:2387–2392. - PubMed
-
- Ellis C, Moran M, McCormick F, Pawson T. Phosphorylation of GAP and GAP-associated proteins by transforming and mitogenic tyrosine kinases. Nature. 1990;343:377–381. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

