A method for direct measurement of protein stability in vivo
- PMID: 19157083
- PMCID: PMC2892803
- DOI: 10.1007/978-1-59745-367-7_7
A method for direct measurement of protein stability in vivo
Abstract
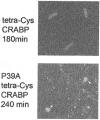
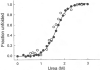
The stability of proteins is tuned by evolution to enable them to perform their cellular functions for the success of an organism. Yet, most of the arsenal of biophysical techniques at our disposal to characterize the thermodynamic stability of proteins is limited to in vitro samples. We describe an approach that we have developed to observe a protein directly in a cell and to monitor a fluorescence signal that reports the unfolding transition of the protein, yielding quantitatively interpretable stability data in vivo. The method is based on incorporation of structurally nonperturbing, specific binding motifs for a bis-arsenical fluorescein derivative in sites that result in dye fluorescence differences between the folded and unfolded states of the protein under study. This fluorescence labeling approach makes possible the determination of thermodynamic stability by direct urea titration in Escherichia coli cells. The specific case study we describe was carried out on the predominantly beta-sheet intracellular lipid-binding protein, cellular retinoic acid-binding protein (CRABP), expressed in E. coli.
Figures
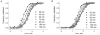

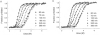
Similar articles
-
Chapter 3: A fluorescent window into protein folding and aggregation in cells.Methods Cell Biol. 2008;89:59-70. doi: 10.1016/S0091-679X(08)00603-1. Methods Cell Biol. 2008. PMID: 19118672 Free PMC article.
-
Multistate equilibrium unfolding of Escherichia coli dihydrofolate reductase: thermodynamic and spectroscopic description of the native, intermediate, and unfolded ensembles.Biochemistry. 2000 Aug 8;39(31):9540-50. doi: 10.1021/bi000511y. Biochemistry. 2000. PMID: 10924151
-
Selective stabilization of a partially unfolded protein by a metabolite.J Mol Biol. 2012 Sep 21;422(3):403-13. doi: 10.1016/j.jmb.2012.05.044. Epub 2012 Jun 7. J Mol Biol. 2012. PMID: 22684147
-
Engineering more stable proteins.Chem Soc Rev. 2018 Dec 21;47(24):9026-9045. doi: 10.1039/c8cs00014j. Epub 2018 Oct 11. Chem Soc Rev. 2018. PMID: 30306986 Review.
-
Thermodynamics of protein folding: effects of hydration and electrostatic interactions.Adv Biophys. 1994;30:105-54. doi: 10.1016/0065-227x(94)90012-4. Adv Biophys. 1994. PMID: 7709803 Review.
Cited by
-
Protein stability [determination] problems.Front Mol Biosci. 2022 Aug 5;9:880358. doi: 10.3389/fmolb.2022.880358. eCollection 2022. Front Mol Biosci. 2022. PMID: 35992266 Free PMC article.
-
Quantifying the thermodynamics of protein unfolding using 2D NMR spectroscopy.Commun Chem. 2020 Aug 7;3:100. doi: 10.1038/s42004-020-00358-1. Commun Chem. 2020. PMID: 33718626 Free PMC article.
-
Determining biophysical protein stability in lysates by a fast proteolysis assay, FASTpp.PLoS One. 2012;7(10):e46147. doi: 10.1371/journal.pone.0046147. Epub 2012 Oct 3. PLoS One. 2012. PMID: 23056252 Free PMC article.
References
-
- Chiti F, Stefani M, Taddei N, et al. Rationalization of the effects of mutations on peptide and protein aggrcgation rates. Nature. 2003;424:805–808. - PubMed
-
- Dobson CM. Protein folding and misfolding. Nature. 2003;426:884–890. - PubMed
-
- Evans MS, Clarke TF, IV, Clark PL. Conformations of co-translational folding intermediates. Protein Pept Lett. 2005;12:189–195. - PubMed
-
- Minton AP. How can biochemical reactions within cells differ from those in test tubes? J Cell Sci. 2006;119:2863–2869. - PubMed
-
- Pace CN, McGrath T. Substrate stabilization of lysozyme to thermal and guanidine hydrochloride denaturation. J Biol Chem. 1980;255:3862–3865. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

