Periodic reversal of direction allows Myxobacteria to swarm
- PMID: 19164578
- PMCID: PMC2633568
- DOI: 10.1073/pnas.0811662106
Periodic reversal of direction allows Myxobacteria to swarm
Abstract
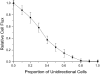
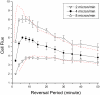
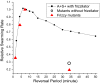
Many bacteria can rapidly traverse surfaces from which they are extracting nutrient for growth. They generate flat, spreading colonies, called swarms because they resemble swarms of insects. We seek to understand how members of any dense swarm spread efficiently while being able to perceive and interfere minimally with the motion of others. To this end, we investigate swarms of the myxobacterium, Myxococcus xanthus. Individual M. xanthus cells are elongated; they always move in the direction of their long axis; and they are in constant motion, repeatedly touching each other. Remarkably, they regularly reverse their gliding directions. We have constructed a detailed cell- and behavior-based computational model of M. xanthus swarming that allows the organization of cells to be computed. By using the model, we are able to show that reversals of gliding direction are essential for swarming and that reversals increase the outflow of cells across the edge of the swarm. Cells at the swarm edge gain maximum exposure to nutrient and oxygen. We also find that the reversal period predicted to maximize the outflow of cells is the same (within the errors of measurement) as the period observed in experiments with normal M. xanthus cells. This coincidence suggests that the circuit regulating reversals evolved to its current sensitivity under selection for growth achieved by swarming. Finally, we observe that, with time, reversals increase the cell alignment, and generate clusters of parallel cells.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



References
-
- Harshey RM. Bees aren't the only ones: Swarming in Gram-negative bacteria. Mol Microbiol. 1994;13:389–394. - PubMed
-
- Warren JW. Catheter-associated urinary tract infection. In: Bayless T, Brain M, Cherniak R, editors. Current Therapy in Internal Medicine. Philadelphia: BC Decker Inc.; 1987. pp. 290–292.
-
- Helbing D. Traffic and related self-driven many-particle systems. Rev Mod Phys. 2001;73:1067–1141.
-
- McCarter LL. Dual flagellar systems enable motility under different circumstances. J Mol Microbiol Biotechnol. 2004;7:18–29. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

