Positive fluorescent selection permits precise, rapid, and in-depth overexpression analysis in plant protoplasts
- PMID: 19168642
- PMCID: PMC2649414
- DOI: 10.1104/pp.108.133975
Positive fluorescent selection permits precise, rapid, and in-depth overexpression analysis in plant protoplasts
Erratum in
- Plant Physiol. 2009 Jun;150(2):1105
Abstract
Transient genetic modification of plant protoplasts is a straightforward and rapid technique for the study of numerous aspects of plant biology. Recent studies in metazoan systems have utilized cell-based assays to interrogate signal transduction pathways using high-throughput methods. Plant biologists could benefit from new tools that expand the use of cell culture for large-scale analysis of gene function. We have developed a system that employs fluorescent positive selection in combination with flow cytometric analysis and fluorescence-activated cell sorting to isolate responses in the transformed protoplasts exclusively. The system overcomes the drawback that transfected protoplast suspensions are often a heterogeneous mix of cells that have and have not been successfully transformed. This Gateway-compatible system enables high-throughput screening of genetic circuitry using overexpression. The incorporation of a red fluorescent protein selection marker enables combined utilization with widely available green fluorescent protein (GFP) tools. For instance, such a dual labeling approach allows cytometric analysis of GFP reporter gene activation expressly in the transformed cells or fluorescence-activated cell sorting-mediated isolation and downstream examination of overexpression effects in a specific GFP-marked cell population. Here, as an example, novel uses of this system are applied to the study of auxin signaling, exploiting the red fluorescent protein/GFP dual labeling capability. In response to manipulation of the auxin response network through overexpression of dominant negative auxin signaling components, we quantify effects on auxin-responsive DR5::GFP reporter gene activation as well as profile genome-wide transcriptional changes specifically in cells expressing a root epidermal marker.
Figures



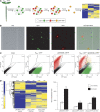
References
-
- Birnbaum K, Shasha DE, Wang JY, Jung JW, Lambert GM, Galbraith DW, Benfey PN (2003) A gene expression map of the Arabidopsis root. Science 302 1956–1960 - PubMed
-
- Brady SM, Orlando DA, Lee JY, Wang JY, Koch J, Dinneny JR, Mace D, Ohler U, Benfey PN (2007) A high-resolution root spatiotemporal map reveals dominant expression patterns. Science 318 801–806 - PubMed
-
- Cummins I, Steel PG, Edwards R (2007) Identification of a carboxylesterase expressed in protoplasts using fluorescence-activated cell sorting. Plant Biotechnol J 5 354–359 - PubMed
-
- De Sutter V, Vanderhaeghen R, Tilleman S, Lammertyn F, Vanhoutte I, Karimi M, Inzé D, Goossens A, Hilson P (2005) Exploration of jasmonate signalling via automated and standardized transient expression assays in tobacco cells. Plant J 44 1065–1076 - PubMed
-
- Dinneny JR, Long TA, Wang JY, Jung JW, Mace D, Pointer S, Barron C, Brady SM, Schiefelbein J, Benfey PN (2008) Cell identity mediates the response of Arabidopsis roots to abiotic stress. Science 320 942–945 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

