Viewing the bilayer hydrocarbon core using neutron diffraction
- PMID: 19169614
- PMCID: PMC2667903
- DOI: 10.1007/s00232-008-9151-3
Viewing the bilayer hydrocarbon core using neutron diffraction
Abstract
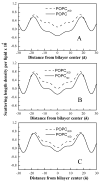
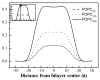
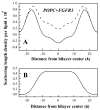
Membrane proteins fold, assemble and function within their native fluid lipid environment. Structural studies of fluid lipid bilayers are thus critically important for understanding processes in membranes. Here, we propose a simple approach to visualize the hydrocarbon core using neutron diffraction and deuterated lipids that are commercially available. This method should have broad utility in structural studies of the bilayer response to protein insertion and folding in membranes.
Figures


Similar articles
-
The structure of polyunsaturated lipid bilayers important for rhodopsin function: a neutron diffraction study.Biophys J. 2006 Jan 1;90(1):L04-6. doi: 10.1529/biophysj.105.071712. Epub 2005 Oct 28. Biophys J. 2006. PMID: 16258049 Free PMC article.
-
The impact of deuteration on natural and synthetic lipids: A neutron diffraction study.Colloids Surf B Biointerfaces. 2018 Aug 1;168:126-133. doi: 10.1016/j.colsurfb.2018.02.009. Epub 2018 Feb 5. Colloids Surf B Biointerfaces. 2018. PMID: 29433911
-
Efficient internalization of TAT peptide in zwitterionic DOPC phospholipid membrane revealed by neutron diffraction.Biochim Biophys Acta Biomembr. 2017 May;1859(5):910-916. doi: 10.1016/j.bbamem.2017.01.036. Epub 2017 Jan 31. Biochim Biophys Acta Biomembr. 2017. PMID: 28153495
-
Biomolecular and amphiphilic films probed by surface sensitive X-ray and neutron scattering.Anal Bioanal Chem. 2004 Aug;379(7-8):960-73. doi: 10.1007/s00216-004-2696-9. Epub 2004 Jul 31. Anal Bioanal Chem. 2004. PMID: 15338090 Review.
-
Multiscale lipid membrane dynamics as revealed by neutron spectroscopy.Prog Lipid Res. 2022 Jul;87:101179. doi: 10.1016/j.plipres.2022.101179. Epub 2022 Jun 30. Prog Lipid Res. 2022. PMID: 35780913 Review.
Cited by
-
A membrane-translocating peptide penetrates into bilayers without significant bilayer perturbations.Biophys J. 2013 Jun 4;104(11):2419-28. doi: 10.1016/j.bpj.2013.04.043. Biophys J. 2013. PMID: 23746514 Free PMC article.
-
Neutron diffraction studies of the interaction between amphotericin B and lipid-sterol model membranes.Sci Rep. 2012;2:778. doi: 10.1038/srep00778. Epub 2012 Oct 29. Sci Rep. 2012. PMID: 23110248 Free PMC article.
References
-
- White SH, Wimley WC, Ladokhin AS, Hristova K. Protein folding in membranes: Pondering the nature of the bilayer milieu. Biol Skr Dan Selsk. 1998;49:91–98.
-
- White SH, Ladokhin AS, Jayasinghe S, Hristova K. How membranes shape protein structure. J Biol Chem. 2001;276:32395–32398. - PubMed
-
- White SH, Wimley WC. Membrane protein folding and stability: Physical principles. Annu Rev Biophys Biomol Struc. 1999;28:319–365. - PubMed
-
- MacKenzie KR. Folding and stability of alpha-helical integral membrane proteins. Chem Rev. 2006;106:1931–1977. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

