Design of 240,000 orthogonal 25mer DNA barcode probes
- PMID: 19171886
- PMCID: PMC2631075
- DOI: 10.1073/pnas.0812506106
Design of 240,000 orthogonal 25mer DNA barcode probes
Abstract
DNA barcodes linked to genetic features greatly facilitate screening these features in pooled formats using microarray hybridization, and new tools are needed to design large sets of barcodes to allow construction of large barcoded mammalian libraries such as shRNA libraries. Here we report a framework for designing large sets of orthogonal barcode probes. We demonstrate the utility of this framework by designing 240,000 barcode probes and testing their performance by hybridization. From the test hybridizations, we also discovered new probe design rules that significantly reduce cross-hybridization after their introduction into the framework of the algorithm. These rules should improve the performance of DNA microarray probe designs for many applications.
Conflict of interest statement
The authors declare no conflict of interest.
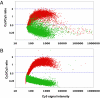
Figures


Similar articles
-
Designing DNA barcodes orthogonal in melting temperature by simulated annealing optimization.Nucleic Acid Ther. 2013 Apr;23(2):140-51. doi: 10.1089/nat.2012.0394. Nucleic Acid Ther. 2013. PMID: 23557118
-
Systematic validation and atomic force microscopy of non-covalent short oligonucleotide barcode microarrays.PLoS One. 2008 Feb 6;3(2):e1546. doi: 10.1371/journal.pone.0001546. PLoS One. 2008. PMID: 18253494 Free PMC article.
-
ProbeMaker: an extensible framework for design of sets of oligonucleotide probes.BMC Bioinformatics. 2005 Sep 19;6:229. doi: 10.1186/1471-2105-6-229. BMC Bioinformatics. 2005. PMID: 16171527 Free PMC article.
-
Real-time assays with molecular beacons and other fluorescent nucleic acid hybridization probes.Clin Chim Acta. 2006 Jan;363(1-2):48-60. doi: 10.1016/j.cccn.2005.04.037. Epub 2005 Aug 18. Clin Chim Acta. 2006. PMID: 16111667 Review.
-
Recent patents and challenges on DNA microarray probe design technologies.Recent Pat DNA Gene Seq. 2010 Nov;4(3):210-7. doi: 10.2174/187221510794751640. Recent Pat DNA Gene Seq. 2010. PMID: 21121887 Review.
Cited by
-
CD8+ T cells from patients with narcolepsy and healthy controls recognize hypocretin neuron-specific antigens.Nat Commun. 2019 Feb 19;10(1):837. doi: 10.1038/s41467-019-08774-1. Nat Commun. 2019. PMID: 30783092 Free PMC article.
-
Spatial transcriptomics using combinatorial fluorescence spectral and lifetime encoding, imaging and analysis.Nat Commun. 2022 Jan 10;13(1):169. doi: 10.1038/s41467-021-27798-0. Nat Commun. 2022. PMID: 35013281 Free PMC article.
-
Evolution of MHC-based technologies used for detection of antigen-responsive T cells.Cancer Immunol Immunother. 2017 May;66(5):657-666. doi: 10.1007/s00262-017-1971-5. Epub 2017 Mar 17. Cancer Immunol Immunother. 2017. PMID: 28314956 Free PMC article. Review.
-
A set of experimentally validated, mutually orthogonal primers for combinatorially specifying genetic components.Synth Biol (Oxf). 2018 Jan 18;3(1):ysx008. doi: 10.1093/synbio/ysx008. eCollection 2018. Synth Biol (Oxf). 2018. PMID: 32995509 Free PMC article.
-
Spatial organization shapes the turnover of a bacterial transcriptome.Elife. 2016 May 20;5:e13065. doi: 10.7554/eLife.13065. Elife. 2016. PMID: 27198188 Free PMC article.
References
-
- Winzeler EA, et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science. 1999;285:901–906. - PubMed
-
- Giaever G, et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature. 2002;418:387–391. - PubMed
-
- Shoemaker DD, et al. Quantitative phenotypic analysis of yeast deletion mutants using a highly parallel molecular bar-coding strategy. Nat Genet. 1996;14:450–456. - PubMed
-
- Silva JM, et al. Second-generation shRNA libraries covering the mouse and human genomes. Nat Genet. 2005;37:1281–1288. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

