A gene catalogue for post-diapause development of an anhydrobiotic arthropod Artemia franciscana
- PMID: 19173719
- PMCID: PMC2649162
- DOI: 10.1186/1471-2164-10-52
A gene catalogue for post-diapause development of an anhydrobiotic arthropod Artemia franciscana
Abstract
Background: Diapause is a reversible state of developmental suspension and found among diverse taxa, from plants to animals, including marsupials and some other mammals. Although previous work has accumulated ample data, the molecular mechanism underlying diapause and reactivation from it remain elusive.
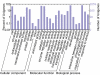
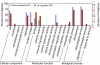
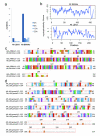
Results: Using Artemia franciscana, a model organism to study the development of post-diapause embryos in Arthropod, we sequenced random clones up to a total of 28,039 ESTs from four cDNA libraries made from dehydrated cysts and three time points after rehydration/reactivation, which were assembled into 8,018 unigene clusters. We identified 324 differentially-expressed genes (DEGs, P < 0.05) based on pairwise comparisons of the four cDNA libraries. We identified a group of genes that are involved in an anti-water-deficit system, including proteases, protease inhibitors, heat shock proteins, and several novel members of the late embryogenesis abundant (LEA) protein family. In addition, we classified most of the up-regulated genes after cyst reactivation into metabolism, biosynthesis, transcription, and translation, and this result is consistent with the rapid development of the embryo. Some of the specific expressions of DEGs were confirmed experimentally based on quantitative real-time PCR.
Conclusion: We found that the first 5-hour period after rehydration is most important for embryonic reactivation of Artemia. As the total number of expressed genes increases significantly, the majority of DEGs were also identified in this period, including a group of water-deficient-induced genes. A group of genes with similar functions have been described in plant seeds; for instance, one of the novel LEA members shares ~70% amino-acid identity with an Arabidopsis EM (embryonic abundant) protein, the closest animal relative to plant LEA families identified thus far. Our findings also suggested that not only nutrition, but also mRNAs are produced and stored during cyst formation to support rapid development after reactivation.
Figures
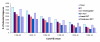
References
-
- Abatzopoulos TJB, J A, Clegg JS, Sorgeloos P. Artemia: Basic and Applied Biology: Springer; 2002. http://www.springer.com/life+sci/zoology/book/978-1-4020-0746-0 - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Research Materials

