Complete genome of the broad-host-range Erwinia amylovora phage phiEa21-4 and its relationship to Salmonella phage felix O1
- PMID: 19181832
- PMCID: PMC2663227
- DOI: 10.1128/AEM.02352-08
Complete genome of the broad-host-range Erwinia amylovora phage phiEa21-4 and its relationship to Salmonella phage felix O1
Abstract
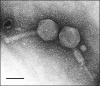
The first complete genome sequence for a myoviridal bacteriophage, PhiEa21-4, infecting Erwinia amylovora, Erwinia pyrifoliae, and Pantoea agglomerans strains has been determined. The unique sequence of this terminally redundant, circularly permuted genome is 84,576 bp. The PhiEa21-4 genome has a GC content of 43.8% and contains 117 putative protein-coding genes and 26 tRNA genes. PhiEa21-4 is the first phage in which a precisely conserved rho-independent terminator has been found dispersed throughout the genome, with 24 copies in all. Also notable in the PhiEa21-4 genome are the presence of tRNAs with six- and nine-base anticodon loops, the absence of a small packaging terminase subunit, and the presence of nadV, a principle component of the NAD(+) salvage pathway, which has been found in only a few phage genomes to date. PhiEa21-4 is the first reported Felix O1-like phage genome; 56% of the predicted PhiEa21-4 proteins share homology with those of the Salmonella phage. Apart from this similarity to Felix O1, the PhiEa21-4 genome appears to be substantially different, both globally and locally, from previously reported sequences. A total of 43 of the 117 genes are unique to PhiEa21-4, and 32 of the Felix O1-like genes do not appear in any phage genome sequences other than PhiEa21-4 and Felix O1. N-terminal sequencing and matrix-assisted laser desorption ionization-time of flight analysis resulted in the identification of five PhiEa21-4 genes coding for virion structural proteins, including the major capsid protein.
Figures


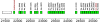
Similar articles
-
Novel virulent and broad-host-range Erwinia amylovora bacteriophages reveal a high degree of mosaicism and a relationship to Enterobacteriaceae phages.Appl Environ Microbiol. 2011 Sep;77(17):5945-54. doi: 10.1128/AEM.03022-10. Epub 2011 Jul 15. Appl Environ Microbiol. 2011. PMID: 21764969 Free PMC article.
-
The genome and proteome of a virulent Escherichia coli O157:H7 bacteriophage closely resembling Salmonella phage Felix O1.Virol J. 2009 Apr 20;6:41. doi: 10.1186/1743-422X-6-41. Virol J. 2009. PMID: 19379502 Free PMC article.
-
Characterization of a new ViI-like Erwinia amylovora bacteriophage phiEa2809.FEMS Microbiol Lett. 2015 Apr;362(7):fnv031. doi: 10.1093/femsle/fnv031. Epub 2015 Feb 24. FEMS Microbiol Lett. 2015. PMID: 25714551
-
A bacteriophage reagent for Salmonella: molecular studies on Felix 01.Int J Food Microbiol. 2002 Apr 5;74(3):217-27. doi: 10.1016/s0168-1605(01)00682-1. Int J Food Microbiol. 2002. PMID: 11981972 Review.
-
The SPO1-related bacteriophages.Arch Virol. 2010 Oct;155(10):1547-61. doi: 10.1007/s00705-010-0783-0. Epub 2010 Aug 17. Arch Virol. 2010. PMID: 20714761 Review.
Cited by
-
CoreGenes3.5: a webserver for the determination of core genes from sets of viral and small bacterial genomes.BMC Res Notes. 2013 Apr 8;6:140. doi: 10.1186/1756-0500-6-140. BMC Res Notes. 2013. PMID: 23566564 Free PMC article.
-
Complete Genome of the Xanthomonas euvesicatoria Specific Bacteriophage KΦ1, Its Survival and Potential in Control of Pepper Bacterial Spot.Front Microbiol. 2018 Aug 29;9:2021. doi: 10.3389/fmicb.2018.02021. eCollection 2018. Front Microbiol. 2018. PMID: 30210484 Free PMC article.
-
Prediction of key amino acids of Salmonella phage endolysin LysST-3 and detection of its mutants' activity.Arch Microbiol. 2024 Mar 11;206(4):151. doi: 10.1007/s00203-024-03915-7. Arch Microbiol. 2024. PMID: 38467842
-
Phage Therapy: What Have We Learned?Viruses. 2018 May 28;10(6):288. doi: 10.3390/v10060288. Viruses. 2018. PMID: 29843391 Free PMC article. Review.
-
The arable ecosystem as battleground for emergence of new human pathogens.Front Microbiol. 2014 Mar 20;5:104. doi: 10.3389/fmicb.2014.00104. eCollection 2014. Front Microbiol. 2014. PMID: 24688484 Free PMC article. Review.
References
-
- Anderson, J. C., T. J. Magliery, and P. G. Schultz. 2002. Exploring the limits of codon and anticodon size. Chem. Biol. 9:237-244. - PubMed
-
- Cherry, W. B., B. R. Davis, P. R. Edwards, and R. B. Hogan. 1954. A simple procedure for the identification of the genus Salmonella by means of a specific bacteriophage. J. Lab. Clin. Med. 44:51-55. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

