Evolutionary history of the phl gene cluster in the plant-associated bacterium Pseudomonas fluorescens
- PMID: 19181839
- PMCID: PMC2663185
- DOI: 10.1128/AEM.02052-08
Evolutionary history of the phl gene cluster in the plant-associated bacterium Pseudomonas fluorescens
Abstract
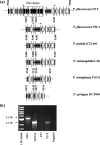
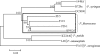
Pseudomonas fluorescens is of agricultural and economic importance as a biological control agent largely because of its plant association and production of secondary metabolites, in particular 2,4-diacetylphloroglucinol (2,4-DAPG). This polyketide, which is encoded by the eight-gene phl cluster, has antimicrobial effects on phytopathogens, promotes amino acid exudation from plant roots, and induces systemic resistance in plants. Despite its importance, 2,4-DAPG production is limited to a subset of P. fluorescens strains. Determination of the evolution of the phl cluster and understanding the selective pressures promoting its retention or loss in lineages of P. fluorescens will help in the development of P. fluorescens as a viable and effective inoculant for application in agriculture. In this study, genomic and sequence-based approaches were integrated to reconstruct the phylogeny of P. fluorescens and the phl cluster. It was determined that 2,4-DAPG production is an ancestral trait in the species P. fluorescens but that most lineages have lost this capacity through evolution. Furthermore, intragenomic recombination has relocated the phl cluster within the P. fluorescens genome at least three times, but the integrity of the cluster has always been maintained. The possible evolutionary and functional implications for retention of the phl cluster and 2,4-DAPG production in some lineages of P. fluorescens are discussed.
Figures



Similar articles
-
Autoinduction of 2,4-diacetylphloroglucinol biosynthesis in the biocontrol agent Pseudomonas fluorescens CHA0 and repression by the bacterial metabolites salicylate and pyoluteorin.J Bacteriol. 2000 Mar;182(5):1215-25. doi: 10.1128/JB.182.5.1215-1225.2000. J Bacteriol. 2000. PMID: 10671440 Free PMC article.
-
Interspecies signaling modulates the biosynthesis of antimicrobial secondary metabolites related to biological control activities of Pseudomonas fluorescens 2P24.Microbiol Spectr. 2025 Mar 4;13(3):e0188624. doi: 10.1128/spectrum.01886-24. Epub 2025 Feb 3. Microbiol Spectr. 2025. PMID: 39898669 Free PMC article.
-
Transcriptional Regulator PhlH Modulates 2,4-Diacetylphloroglucinol Biosynthesis in Response to the Biosynthetic Intermediate and End Product.Appl Environ Microbiol. 2017 Oct 17;83(21):e01419-17. doi: 10.1128/AEM.01419-17. Print 2017 Nov 1. Appl Environ Microbiol. 2017. PMID: 28821548 Free PMC article.
-
Identification and characterization of a gene cluster for synthesis of the polyketide antibiotic 2,4-diacetylphloroglucinol from Pseudomonas fluorescens Q2-87.J Bacteriol. 1999 May;181(10):3155-63. doi: 10.1128/JB.181.10.3155-3163.1999. J Bacteriol. 1999. PMID: 10322017 Free PMC article.
-
Role of 2,4-diacetylphloroglucinol-producing fluorescent Pseudomonas spp. in the defense of plant roots.Plant Biol (Stuttg). 2007 Jan;9(1):4-20. doi: 10.1055/s-2006-924473. Epub 2006 Oct 23. Plant Biol (Stuttg). 2007. PMID: 17058178 Review.
Cited by
-
Phloroglucinol Derivatives in Plant-Beneficial Pseudomonas spp.: Biosynthesis, Regulation, and Functions.Metabolites. 2021 Mar 20;11(3):182. doi: 10.3390/metabo11030182. Metabolites. 2021. PMID: 33804595 Free PMC article. Review.
-
Structure and Catalytic Mechanism of a Bacterial Friedel-Crafts Acylase.Chembiochem. 2019 Jan 2;20(1):88-95. doi: 10.1002/cbic.201800462. Epub 2018 Nov 26. Chembiochem. 2019. PMID: 30318713 Free PMC article.
-
Genome sequence of the biocontrol strain Pseudomonas fluorescens F113.J Bacteriol. 2012 Mar;194(5):1273-4. doi: 10.1128/JB.06601-11. J Bacteriol. 2012. PMID: 22328765 Free PMC article.
-
Antifungal Bacteria on Woodland Salamander Skin Exhibit High Taxonomic Diversity and Geographic Variability.Appl Environ Microbiol. 2017 Apr 17;83(9):e00186-17. doi: 10.1128/AEM.00186-17. Print 2017 May 1. Appl Environ Microbiol. 2017. PMID: 28213545 Free PMC article.
-
Unlocking the potential of ecofriendly guardians for biological control of plant diseases, crop protection and production in sustainable agriculture.3 Biotech. 2025 Apr;15(4):82. doi: 10.1007/s13205-025-04243-3. Epub 2025 Mar 9. 3 Biotech. 2025. PMID: 40071128 Review.
References
-
- Abbas, A., J. E. McGuire, D. Crowley, C. Bayesse, M. Fow, and F. O'Gara. 2004. The putative permease PhlE of Pseudomonas fluorescens F113 has a role in 2,4-diacetylphloroglucinol resistance and in general stress tolerance. Microbiology 150:2443-2450. - PubMed
-
- Abbott, J. C., D. M. Aanensen, K. Rutherford, S. Butcher, and B. G. Spratt. 2005. WebACT—an online companion for the artemis comparison tool. Bioinformatics 21:3665-3666. - PubMed
-
- Achouak, W., L. Sutra, T. Heulin, J. M. Meyer, N. Fromin, S. Degraeve, R. Christen, and L. Gardan. 2000. Pseudomonas brassicacearum sp. nov. and Pseudomonas thivervalensis sp. nov., two root-associated bacteria isolated from Brassica napus and Arabidopsis thaliana. Int. J. Syst. Evol. Microbiol. 50:9-18. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources

