Biological basis for restriction of microRNA targets to the 3' untranslated region in mammalian mRNAs
- PMID: 19182800
- PMCID: PMC2713750
- DOI: 10.1038/nsmb.1552
Biological basis for restriction of microRNA targets to the 3' untranslated region in mammalian mRNAs
Abstract
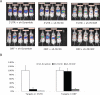
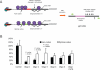
MicroRNAs (miRNAs) interact with target sites located in the 3' untranslated regions (3' UTRs) of mRNAs to downregulate their expression when the appropriate miRNA is bound to target mRNA. To establish the functional importance of target-site localization in the 3' UTR, we modified the stop codon to extend the coding region of the transgene reporter through the miRNA target sequence. As a result, the miRNAs lost their ability to inhibit translation but retained their ability to function as small interfering RNAs in mammalian cells in culture and in vivo. The addition of rare but not optimal codons upstream of the extended opening reading frame (ORF) made the miRNA target site more accessible and restored miRNA-induced translational knockdown. Taken together, these results suggest that active translation impedes miRNA-programmed RISC association with target mRNAs and support a mechanistic explanation for the localization of most miRNA target sites in noncoding regions of mRNAs in mammals.
Figures


References
-
- Ambros V. The functions of animal microRNAs. Nature. 2004;431:350–5. - PubMed
-
- Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–97. - PubMed
-
- Berezikov E, et al. Phylogenetic shadowing and computational identification of human microRNA genes. Cell. 2005;120:21–4. - PubMed
-
- Lewis BP, Burge CB, Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell. 2005;120:15–20. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

