Use of weighted reference panels based on empirical estimates of ancestry for capturing untyped variation
- PMID: 19184111
- PMCID: PMC3126674
- DOI: 10.1007/s00439-009-0627-8
Use of weighted reference panels based on empirical estimates of ancestry for capturing untyped variation
Abstract
Many association methods use a subset of genotyped single nucleotide polymorphisms (SNPs) to capture or infer genotypes at other untyped SNPs. We and others previously showed that tag SNPs selected to capture common variation using data from The International HapMap Consortium (Nature 437:1299-1320, 2005), The International HapMap Consortium (Nature 449:851-861, 2007) could also capture variation in populations of similar ancestry to HapMap reference populations (de Bakker et al. in Nat Genet 38:1298-1303, 2006; González-Neira et al. in Genome Res 16:323-330, 2006; Montpetit et al. in PLoS Genet 2:282-290, 2006; Mueller et al. in Am J Hum Genet 76:387-398, 2005). To capture variation in admixed populations or populations less similar to HapMap panels, a "cosmopolitan approach," in which all samples from HapMap are used as a single reference panel, was proposed. Here we refine this suggestion and show that use of a "weighted reference panel," constructed based on empirical estimates of ancestry in the target population (relative to available reference panels), is more efficient than the cosmopolitan approach. Weighted reference panels capture, on average, only slightly fewer common variants (minor allele frequency > 5%) than the cosmopolitan approach (mean r (2) = 0.977 vs. 0.989, 94.5% variation captured vs. 96.8% at r (2) > 0.8), across the five populations of the Multiethnic Cohort, but entail approximately 25% fewer tag SNPs per panel (average 538 vs. 718). These results extend a recent study in two Indian populations (Pemberton et al. in Ann Hum Genet 72:535-546, 2008). Weighted reference panels are potentially useful for both the selection of tag SNPs in diverse populations and perhaps in the design of reference panels for imputation of untyped genotypes in genome-wide association studies in admixed populations.
Figures
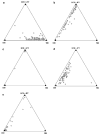
References
-
- Barrett JC, Fry B, Maller J, Daly MJ. Haploview: analysis and visualization of ld and haplotype maps. Bioinformatics. 2005;21:263–265. - PubMed
-
- Cann HM, de Toma C, Cazes L, Legrand MF, Morel V, Piouffre L, Bodmer J, Bodmer WF, Bonne-Tamir B, Cambon-Thomsen A, Chen Z, Chu J, Carcassi C, Contu L, Du R, Excoffier L, Ferrara GB, Friedlaender JS, Groot H, Gurwitz D, Jenkins T, Herrera RJ, Huang X, Kidd J, Kidd KK, Langaney A, Lin AA, Mehdi SQ, Parham P, Piazza A, Pistillo MP, Qian Y, Shu Q, Xu J, Zhu S, Weber JL, Greely HT, Feldman MW, Thomas G, Dausset J, Cavalli-Sforza LL. A human genome diversity cell line panel. Science. 2002;296:261–262. - PubMed
-
- Chapman JM, Cooper JD, Todd JA, Clayton DG. Detecting disease associations due to linkage disequilibrium using haplotype tags: a class of tests and determinants of statistical power. Hum Hered. 2003;56:18–31. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

