Dictyostelium cells bind a secreted autocrine factor that represses cell proliferation
- PMID: 19187549
- PMCID: PMC2644720
- DOI: 10.1186/1471-2091-10-4
Dictyostelium cells bind a secreted autocrine factor that represses cell proliferation
Abstract
Background: Dictyostelium cells secrete the proteins AprA and CfaD. Cells lacking either AprA or CfaD proliferate faster than wild type, while AprA or CfaD overexpressor cells proliferate slowly, indicating that AprA and CfaD are autocrine factors that repress proliferation. CfaD interacts with AprA and requires the presence of AprA to slow proliferation. To determine if CfaD is necessary for the ability of AprA to slow proliferation, whether AprA binds to cells, and if so whether the binding requires the presence of CfaD, we examined the binding and effect on proliferation of recombinant AprA.
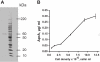
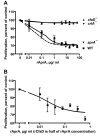
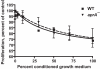
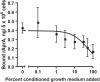
Results: We find that the extracellular accumulation of AprA increases with cell density and reaches a concentration of 0.3 microg/ml near a stationary cell density. When added to wild-type or aprA- cells, recombinant AprA (rAprA) significantly slows proliferation at 0.1 microg/ml and higher concentrations. From 4 to 64 microg/ml, the effect of rAprA is at a plateau, slowing but not stopping proliferation. The proliferation-inhibiting activity of rAprA is roughly the same as that of native AprA in conditioned growth medium. Proliferating aprA- cells show saturable binding of rAprA to 92,000 +/- 11,000 cell-surface receptors with a KD of 0.03 +/- 0.02 microg/ml. There appears to be one class of binding site, and no apparent cooperativity. Native AprA inhibits the binding of rAprA to aprA- cells with a Ki of 0.03 mug/ml, suggesting that the binding kinetics of rAprA are similar to those of native AprA. The proliferation of cells lacking CrlA, a cAMP receptor-like protein, or cells lacking CfaD are not affected by rAprA. Surprisingly, both cell types still bind rAprA.
Conclusion: Together, the data suggest that AprA functions as an autocrine proliferation-inhibiting factor by binding to cell surface receptors. Although AprA requires CfaD for activity, it does not require CfaD to bind to cells, suggesting the possibility that cells have an AprA receptor and a CfaD receptor, and activation of both receptors is required to slow proliferation. We previously found that crlA- cells are sensitive to CfaD. Combined with the results presented here, this suggests that CrlA is not the AprA or CfaD receptor, and may be the receptor for an unknown third factor that is required for AprA and CfaD activity.
Figures



Similar articles
-
A Dictyostelium chalone uses G proteins to regulate proliferation.BMC Biol. 2009 Jul 27;7:44. doi: 10.1186/1741-7007-7-44. BMC Biol. 2009. PMID: 19635129 Free PMC article.
-
A Dictyostelium secreted factor requires a PTEN-like phosphatase to slow proliferation and induce chemorepulsion.PLoS One. 2013;8(3):e59365. doi: 10.1371/journal.pone.0059365. Epub 2013 Mar 12. PLoS One. 2013. PMID: 23555023 Free PMC article.
-
The putative bZIP transcription factor BzpN slows proliferation and functions in the regulation of cell density by autocrine signals in Dictyostelium.PLoS One. 2011;6(7):e21765. doi: 10.1371/journal.pone.0021765. Epub 2011 Jul 7. PLoS One. 2011. PMID: 21760904 Free PMC article.
-
Cell density sensing and size determination.Dev Growth Differ. 2011 May;53(4):482-94. doi: 10.1111/j.1440-169X.2010.01248.x. Epub 2011 Apr 27. Dev Growth Differ. 2011. PMID: 21521184 Free PMC article. Review.
-
PSF and CMF, autocrine factors that regulate gene expression during growth and early development of Dictyostelium.Experientia. 1995 Dec 18;51(12):1124-34. doi: 10.1007/BF01944730. Experientia. 1995. PMID: 8536800 Review.
Cited by
-
A Dictyostelium chalone uses G proteins to regulate proliferation.BMC Biol. 2009 Jul 27;7:44. doi: 10.1186/1741-7007-7-44. BMC Biol. 2009. PMID: 19635129 Free PMC article.
-
PTEN and the PTEN-like phosphatase CnrN have both distinct and overlapping roles in a Dictyostelium chemorepulsion pathway.J Cell Sci. 2024 Aug 1;137(15):jcs262054. doi: 10.1242/jcs.262054. Epub 2024 Aug 1. J Cell Sci. 2024. PMID: 38940195 Free PMC article.
-
An altered transcriptome underlies cln5-deficiency phenotypes in Dictyostelium discoideum.Front Genet. 2022 Nov 10;13:1045738. doi: 10.3389/fgene.2022.1045738. eCollection 2022. Front Genet. 2022. PMID: 36437924 Free PMC article.
-
The p21-activated kinase (PAK) family member PakD is required for chemorepulsion and proliferation inhibition by autocrine signals in Dictyostelium discoideum.PLoS One. 2014 May 5;9(5):e96633. doi: 10.1371/journal.pone.0096633. eCollection 2014. PLoS One. 2014. PMID: 24797076 Free PMC article.
-
α/β Hydrolases: Toward Unraveling Entangled Classification.Proteins. 2025 Apr;93(4):855-870. doi: 10.1002/prot.26776. Epub 2024 Dec 2. Proteins. 2025. PMID: 39623291 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

