A toc159 import receptor mutant, defective in hydrolysis of GTP, supports preprotein import into chloroplasts
- PMID: 19188370
- PMCID: PMC2659226
- DOI: 10.1074/jbc.M804235200
A toc159 import receptor mutant, defective in hydrolysis of GTP, supports preprotein import into chloroplasts
Abstract
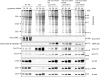
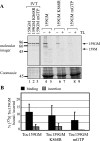
The heterotrimeric Toc core complex of the chloroplast protein import apparatus contains two GTPases, Toc159 and Toc34, together with the protein-conducting channel Toc75. Toc159 and Toc34 are exposed at the chloroplast surface and function in preprotein recognition. Together, they have been shown to facilitate the import of photosynthetic proteins into chloroplasts in Arabidopsis. Consequently, the ppi2 mutant lacking atToc159 has a non-photosynthetic albino phenotype. Previous mutations in the conserved G1 and G3 GTPase motifs abolished the function of Toc159 in vivo by disrupting targeting of the receptor to chloroplasts. Here, we demonstrate that a mutant in a conserved G1 lysine (atToc159 K868R) defective in GTP binding and hydrolysis can target and assemble into Toc complexes. We show that atToc159 K868R can support protein import into isolated chloroplasts, albeit at lower preprotein binding and import efficiencies compared with the wild-type receptor. Considering the absence of measurable GTPase activity in the K868R mutant, we conclude that GTP hydrolysis at atToc159 is not strictly required for preprotein translocation. The data also indicate that preprotein import requires at least one additional GTPase other than Toc159.
Figures




References
-
- Bruce, B. D. (2000) Trends Cell Biol. 10 440-447 - PubMed
-
- Richter, S., and Lamppa, G. K. (2003) J. Biol. Chem. 278 39497-39502 - PubMed
-
- Oreb, M., Tews, I., and Schleiff, E. (2008) Trends Cell Biol. 18 19-27 - PubMed
-
- Kessler, F., and Schnell, D. J. (2006) Traffic 7 248-257 - PubMed
-
- Jarvis, P. (2008) New Phytol. 179 257-285 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

