Forward RNAi screens in primary human hematopoietic stem/progenitor cells
- PMID: 19188664
- PMCID: PMC2670788
- DOI: 10.1182/blood-2008-10-176396
Forward RNAi screens in primary human hematopoietic stem/progenitor cells
Abstract
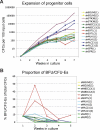
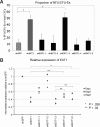
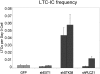
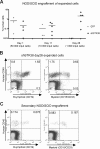
The mechanisms regulating key fate decisions such as self-renewal and differentiation in hematopoietic stem and progenitor cells (HSPC) remain poorly understood. We report here a screening strategy developed to assess modulators of human hematopoiesis using a lentiviral short hairpin RNA (shRNA) library transduced into cord blood-derived stem/progenitor cells. To screen for modifiers of self-renewal/differentiation, we used the limited persistence of HSPCs under ex vivo culture conditions as a baseline for functional selection of shRNAs conferring enhanced maintenance or expansion of the stem/progenitor potential. This approach enables complex, pooled screens in large numbers of cells. Functional selection identified novel specific gene targets (exostoses 1) or shRNA constructs capable of altering human hematopoietic progenitor differentiation or stem cell expansion, respectively, thereby demonstrating the potential of this forward screening approach in primary human stem cell populations.
Figures

References
-
- Sorrentino BP. Clinical strategies for expansion of haematopoietic stem cells. Nat Rev Immunol. 2004;4:878–888. - PubMed
-
- Berns K, Hijmans EM, Mullenders J, et al. A large-scale RNAi screen in human cells identifies new components of the p53 pathway. Nature. 2004;428:431–437. - PubMed
-
- Kolfschoten IG, van Leeuwen B, Berns K, et al. A genetic screen identifies PITX1 as a suppressor of RAS activity and tumorigenicity. Cell. 2005;121:849–858. - PubMed
-
- Moffat J, Grueneberg DA, Yang X, et al. A lentiviral RNAi library for human and mouse genes applied to an arrayed viral high-content screen. Cell. 2006;124:1283–1298. - PubMed
-
- Paddison PJ, Silva JM, Conklin DS, et al. A resource for large-scale RNA-interference-based screens in mammals. Nature. 2004;428:427–431. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

