The CoxD protein of Oligotropha carboxidovorans is a predicted AAA+ ATPase chaperone involved in the biogenesis of the CO dehydrogenase [CuSMoO2] cluster
- PMID: 19189964
- PMCID: PMC2666610
- DOI: 10.1074/jbc.M805354200
The CoxD protein of Oligotropha carboxidovorans is a predicted AAA+ ATPase chaperone involved in the biogenesis of the CO dehydrogenase [CuSMoO2] cluster
Abstract
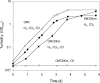
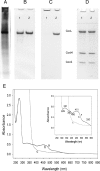
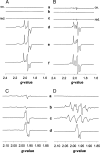
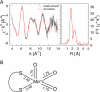
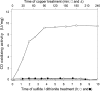
CO dehydrogenase from the Gram-negative chemolithoautotrophic eubacterium Oligotropha carboxidovorans OM5 is a structurally characterized molybdenum-containing iron-sulfur flavoenzyme, which catalyzes the oxidation of CO (CO + H(2)O --> CO(2) + 2e(-) + 2H(+)). It accommodates in its active site a unique bimetallic [CuSMoO(2)] cluster, which is subject to post-translational maturation. Insertional mutagenesis of coxD has established its requirement for the assembly of the [CuSMoO(2)] cluster. Disruption of coxD led to a phenotype of the corresponding mutant OM5 D::km with the following characteristics: (i) It was impaired in the utilization of CO, whereas the utilization of H(2) plus CO(2) was not affected; (ii) Under appropriate induction conditions bacteria synthesized a fully assembled apo-CO dehydrogenase, which could not oxidize CO; (iii) Apo-CO dehydrogenase contained a [MoO(3)] site in place of the [CuSMoO(2)] cluster; and (iv) Employing sodium sulfide first and then the Cu(I)-(thiourea)(3) complex, the non-catalytic [MoO(3)] site could be reconstituted in vitro to a [CuSMoO(2)] cluster capable of oxidizing CO. Sequence information suggests that CoxD is a MoxR-like AAA+ ATPase chaperone related to the hexameric, ring-shaped BchI component of Mg(2+)-chelatases. Recombinant CoxD, which appeared in Escherichia coli in inclusion bodies, occurs exclusively in cytoplasmic membranes of O. carboxidovorans grown in the presence of CO, and its occurrence coincided with GTPase activity upon sucrose density gradient centrifugation of cell extracts. The presumed function of CoxD is the partial unfolding of apo-CO dehydrogenase to assist in the stepwise introduction of sulfur and copper in the [MoO(3)] center of the enzyme.
Figures



References
-
- Auling, G., Busse, J., Hahn, M., Hennecke, H., Kroppenstedt, R., Probst, A., and Stackebrandt, E. (1988) Syst. Appl. Microbiol. 10 264-272
-
- Meyer, O. O. (2005) in Bergey′s Manual of Systematic Bacteriology (Garrity, G. M., ed) pp. 468-471, vol. 2, Springer, New York
-
- Fuhrmann, S., Ferner, M., Jeffke, T., Henne, A., Gottschalk, G., and Meyer, O. (2003) Gene (Amst.) 322 67-75 - PubMed
-
- Santiago, B., Schübel, U., Egelseer, C., and Meyer, O. (1999) Gene (Amst.) 236 115-124 - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

