Cryptic variation in the human mutation rate
- PMID: 19192947
- PMCID: PMC2634788
- DOI: 10.1371/journal.pbio.1000027
Cryptic variation in the human mutation rate
Abstract
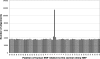
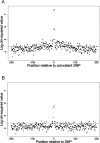
The mutation rate is known to vary between adjacent sites within the human genome as a consequence of context, the most well-studied example being the influence of CpG dinucelotides. We investigated whether there is additional variation by testing whether there is an excess of sites at which both humans and chimpanzees have a single-nucleotide polymorphism (SNP). We found a highly significant excess of such sites, and we demonstrated that this excess is not due to neighbouring nucleotide effects, ancestral polymorphism, or natural selection. We therefore infer that there is cryptic variation in the mutation rate. However, although this variation in the mutation rate is not associated with the adjacent nucleotides, we show that there are highly nonrandom patterns of nucleotides that extend approximately 80 base pairs on either side of sites with coincident SNPs, suggesting that there are extensive and complex context effects. Finally, we estimate the level of variation needed to produce the excess of coincident SNPs and show that there is a similar, or higher, level of variation in the mutation rate associated with this cryptic process than there is associated with adjacent nucleotides, including the CpG effect. We conclude that there is substantial variation in the mutation that has, until now, been hidden from view.
Conflict of interest statement
Competing interests. The authors have declared that no competing interests exist.
Figures
References
-
- Miyata T, Hayashida H, Kuma K, Mitsuyasu K, Yasunaga T. Male-driven molecular evolution: a model and nucleotide sequence analysis. Cold Spring Harb Symp Quant Biol. 1987;52:863–867. - PubMed
-
- Li WH, Yi S, Makova K. Male-driven evolution. Curr Opin Genet Dev. 2002;12:650–656. - PubMed
-
- Lercher MJ, Williams EJ, Hurst LD. Local similarity in evolutionary rates extends over whole chromosomes in human-rodent and mouse-rat comparisons: implications for understanding the mechanistic basis of the male mutation bias. Mol Biol Evol. 2001;18:2032–2039. - PubMed
-
- Matassi G, Sharp PM, Gautier C. Chromosomal location effects on gene sequence evolution in mammals. Curr Biol. 1999;9:786–791. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

