Portrait of an enzyme, a complete structural analysis of a multimodular {beta}-N-acetylglucosaminidase from Clostridium perfringens
- PMID: 19193644
- PMCID: PMC2665110
- DOI: 10.1074/jbc.M808954200
Portrait of an enzyme, a complete structural analysis of a multimodular {beta}-N-acetylglucosaminidase from Clostridium perfringens
Abstract
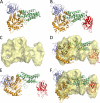
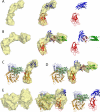
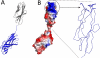
Common features of the extracellular carbohydrate-active virulence factors involved in host-pathogen interactions are their large sizes and modular complexities. This has made them recalcitrant to structural analysis, and therefore our understanding of the significance of modularity in these important proteins is lagging. Clostridium perfringens is a prevalent human pathogen that harbors a wide array of large, extracellular carbohydrate-active enzymes and is an excellent and relevant model system to approach this problem. Here we describe the complete structure of C. perfringens GH84C (NagJ), a 1001-amino acid multimodular homolog of the C. perfringens micro-toxin, which was determined using a combination of small angle x-ray scattering and x-ray crystallography. The resulting structure reveals unprecedented insight into how catalysis, carbohydrate-specific adherence, and the formation of molecular complexes with other enzymes via an ultra-tight protein-protein interaction are spatially coordinated in an enzyme involved in a host-pathogen interaction.
Figures


Similar articles
-
Structural analysis of a bacterial exo-α-D-N-acetylglucosaminidase in complex with an unusual disaccharide found in class III mucin.Glycobiology. 2012 May;22(5):590-5. doi: 10.1093/glycob/cwr165. Epub 2011 Nov 16. Glycobiology. 2012. PMID: 22090394 Free PMC article.
-
Carbohydrate recognition by an architecturally complex α-N-acetylglucosaminidase from Clostridium perfringens.PLoS One. 2012;7(3):e33524. doi: 10.1371/journal.pone.0033524. Epub 2012 Mar 27. PLoS One. 2012. PMID: 22479408 Free PMC article.
-
Probing the Complex Architecture of Multimodular Carbohydrate-Active Enzymes Using a Combination of Small Angle X-Ray Scattering and X-Ray Crystallography.Methods Mol Biol. 2017;1588:239-253. doi: 10.1007/978-1-4939-6899-2_19. Methods Mol Biol. 2017. PMID: 28417374
-
Sialidases From Clostridium perfringens and Their Inhibitors.Front Cell Infect Microbiol. 2020 Jan 10;9:462. doi: 10.3389/fcimb.2019.00462. eCollection 2019. Front Cell Infect Microbiol. 2020. PMID: 31998664 Free PMC article. Review.
-
The interaction of Clostridium perfringens enterotoxin with receptor claudins.Anaerobe. 2016 Oct;41:18-26. doi: 10.1016/j.anaerobe.2016.04.011. Epub 2016 Apr 16. Anaerobe. 2016. PMID: 27090847 Free PMC article. Review.
Cited by
-
Identification and structural analysis of a carbohydrate-binding module specific to alginate, a representative of a new family, CBM96.J Biol Chem. 2023 Feb;299(2):102854. doi: 10.1016/j.jbc.2022.102854. Epub 2022 Dec 31. J Biol Chem. 2023. PMID: 36592931 Free PMC article.
-
A proteomic and cellular analysis of uropods in the pathogen Entamoeba histolytica.PLoS Negl Trop Dis. 2011 Apr 5;5(4):e1002. doi: 10.1371/journal.pntd.0001002. PLoS Negl Trop Dis. 2011. PMID: 21483708 Free PMC article.
-
Rethinking the role of alpha toxin in Clostridium perfringens-associated enteric diseases: a review on bovine necro-haemorrhagic enteritis.Vet Res. 2017 Feb 16;48(1):9. doi: 10.1186/s13567-017-0413-x. Vet Res. 2017. PMID: 28209206 Free PMC article. Review.
-
Structural analysis of a bacterial exo-α-D-N-acetylglucosaminidase in complex with an unusual disaccharide found in class III mucin.Glycobiology. 2012 May;22(5):590-5. doi: 10.1093/glycob/cwr165. Epub 2011 Nov 16. Glycobiology. 2012. PMID: 22090394 Free PMC article.
-
Structural insights of the enzymes from the chitin utilization locus of Flavobacterium johnsoniae.Sci Rep. 2020 Aug 13;10(1):13775. doi: 10.1038/s41598-020-70749-w. Sci Rep. 2020. PMID: 32792608 Free PMC article.
References
-
- Bryant, A. E., Bayer, C. R., Chen, R. Y., Guth, P. H., Wallace, R. J., and Stevens, D. L. (2005) J. Infect. Dis. 192 1014–1022 - PubMed
-
- Hynes, W. (2004) Front. Biosci. 9 3399–3433 - PubMed
-
- Smedley, J. G., 3rd, Fisher, D. J., Sayeed, S., Chakrabarti, G., and McClane, B. A. (2004) Rev. Physiol. Biochem. Pharmacol. 152 183–204 - PubMed
-
- Stevens, D. L., and Bryant, A. E. (2002) Clin. Infect. Dis. 35 S93–S100 - PubMed
-
- Canard, B., Garnier, T., Saint-Joanis, B., and Cole, S. T. (1994) Mol. Gen. Genet. 243 215–224 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

