Characterization of a human immunodeficiency virus type 1 V3 deletion mutation that confers resistance to CCR5 inhibitors and the ability to use aplaviroc-bound receptor
- PMID: 19193800
- PMCID: PMC2663279
- DOI: 10.1128/JVI.01751-08
Characterization of a human immunodeficiency virus type 1 V3 deletion mutation that confers resistance to CCR5 inhibitors and the ability to use aplaviroc-bound receptor
Abstract
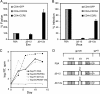
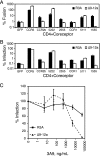
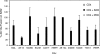
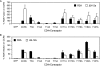
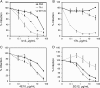
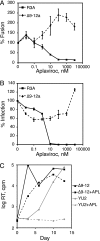
The human immunodeficiency virus type 1 (HIV-1) V3 loop is essential for coreceptor binding and principally determines tropism for the CCR5 and CXCR4 coreceptors. Using the dual-tropic virus HIV-1(R3A), we previously made an extensive panel of V3 deletions and identified subdomains within V3 that could differentially mediate R5 and X4 tropism. A deletion of residues 9 to 12 on the N-terminal side of the V3 stem ablated X4 tropism while leaving R5 tropism intact. This mutation also resulted in complete resistance to several small-molecule CCR5 inhibitors. Here, we extend these studies to further characterize a variant of this mutant, Delta9-12a, adapted for growth in CCR5(+) SupT1 cells. Studies using coreceptor chimeras, monoclonal antibodies directed against the CCR5 amino terminus (NT) and extracellular loops, and CCR5 point mutants revealed that, relative to parental R3A, R5-tropic Delta9-12a was more dependent on the CCR5 NT, a region that contacts the gp120 bridging sheet and V3 base. Neutralization sensitivity assays showed that, compared to parental R3A, Delta9-12a was more sensitive to monoclonal antibodies b12, 4E10, and 2G12. Finally, cross-antagonism assays showed that Delta9-12a could use aplaviroc-bound CCR5 for entry. These studies indicate that increased dependence on the CCR5 NT represents a mechanism by which HIV envelopes acquire resistance to CCR5 antagonists and may have more general implications for mechanisms of drug resistance that arise in vivo. In addition, envelopes such as Delta9-12a may be useful for developing new entry inhibitors that target the interaction of gp120 and the CCR5 NT.
Figures

Similar articles
-
Effects of partial deletions within the human immunodeficiency virus type 1 V3 loop on coreceptor tropism and sensitivity to entry inhibitors.J Virol. 2008 Jan;82(2):664-73. doi: 10.1128/JVI.01793-07. Epub 2007 Oct 31. J Virol. 2008. PMID: 17977968 Free PMC article.
-
V3 loop truncations in HIV-1 envelope impart resistance to coreceptor inhibitors and enhanced sensitivity to neutralizing antibodies.PLoS Pathog. 2007 Aug 24;3(8):e117. doi: 10.1371/journal.ppat.0030117. PLoS Pathog. 2007. PMID: 17722977 Free PMC article.
-
Chemokine (C-C motif) receptor 5-using envelopes predominate in dual/mixed-tropic HIV from the plasma of drug-naive individuals.AIDS. 2008 Jul 31;22(12):1425-31. doi: 10.1097/QAD.0b013e32830184ba. AIDS. 2008. PMID: 18614865
-
[Viral entry as therapeutic target. Current situation of entry inhibitors].Enferm Infecc Microbiol Clin. 2008 Oct;26 Suppl 11:5-11. doi: 10.1016/s0213-005x(08)76557-1. Enferm Infecc Microbiol Clin. 2008. PMID: 19133215 Review. Spanish.
-
Effect of HIV-1 subtype and tropism on treatment with chemokine coreceptor entry inhibitors; overview of viral entry inhibition.Crit Rev Microbiol. 2015;41(4):473-87. doi: 10.3109/1040841X.2013.867829. Epub 2014 Mar 17. Crit Rev Microbiol. 2015. PMID: 24635642 Review.
Cited by
-
Escape from human immunodeficiency virus type 1 (HIV-1) entry inhibitors.Viruses. 2012 Dec;4(12):3859-911. doi: 10.3390/v4123859. Viruses. 2012. PMID: 23342377 Free PMC article. Review.
-
Genomic Evidence for the Nonpathogenic State in HIV-1-Infected Northern Pig-Tailed Macaques.Mol Biol Evol. 2023 May 2;40(5):msad101. doi: 10.1093/molbev/msad101. Mol Biol Evol. 2023. PMID: 37134013 Free PMC article.
-
Design and characterization of a peptide mimotope of the HIV-1 gp120 bridging sheet.Int J Mol Sci. 2012;13(5):5674-5699. doi: 10.3390/ijms13055674. Epub 2012 May 10. Int J Mol Sci. 2012. PMID: 22754323 Free PMC article.
-
Structure-function analysis of human immunodeficiency virus type 1 gp120 amino acid mutations associated with resistance to the CCR5 coreceptor antagonist vicriviroc.J Virol. 2009 Dec;83(23):12151-63. doi: 10.1128/JVI.01351-09. Epub 2009 Sep 23. J Virol. 2009. PMID: 19776131 Free PMC article.
-
CCR5 antibodies HGS004 and HGS101 preferentially inhibit drug-bound CCR5 infection and restore drug sensitivity of Maraviroc-resistant HIV-1 in primary cells.Virology. 2011 Mar 1;411(1):32-40. doi: 10.1016/j.virol.2010.12.029. Epub 2011 Jan 12. Virology. 2011. PMID: 21232779 Free PMC article.
References
-
- Alkhatib, G., C. Combadiere, C. C. Broder, Y. Feng, P. E. Kennedy, P. M. Murphy, and E. A. Berger. 1996. CC CKR5: a RANTES, MIP-1α, MIP-1β receptor as a fusion cofactor for macrophage-tropic HIV-1. Science 2721955-1958. - PubMed
-
- Billick, E., C. Seibert, P. Pugagh, T. Ketas, A. Trkola, M. J. Endres, N. J. Murgolo, E. Coates, G. R. Reyes, B. M. Baroudy, T. P. Sakmar, J. P. Moore, and S. E. Kuhmann. 2004. The differential sensitivity of human and rhesus macaque CCR5 to small-molecule inhibitors of human immunodeficiency virus type 1 entry is explained by a single amino acid difference and suggests a mechanism of action for these inhibitors. J. Virol. 784134-4144. - PMC - PubMed
-
- Braibant, M., S. Brunet, D. Costagliola, C. Rouzioux, H. Agut, H. Katinger, B. Autran, and F. Barin. 2006. Antibodies to conserved epitopes of the HIV-1 envelope in sera from long-term non-progressors: prevalence and association with neutralizing activity. AIDS 201923-1930. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

