MicroRNA-155 modulates the interleukin-1 signaling pathway in activated human monocyte-derived dendritic cells
- PMID: 19193853
- PMCID: PMC2650335
- DOI: 10.1073/pnas.0811073106
MicroRNA-155 modulates the interleukin-1 signaling pathway in activated human monocyte-derived dendritic cells
Abstract
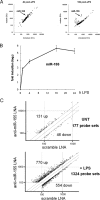
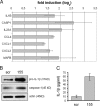
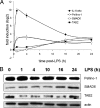
In response to inflammatory stimulation, dendritic cells (DCs) have a remarkable pattern of differentiation (maturation) that exhibits specific mechanisms to control immunity. Here, we show that in response to Lipopolysaccharides (LPS), several microRNAs (miRNAs) are regulated in human monocyte-derived dendritic cells. Among these miRNAs, miR-155 is highly up-regulated during maturation. Using LNA silencing combined to microarray technology, we have identified the Toll-like receptor/interleukin-1 (TLR/IL-1) inflammatory pathway as a general target of miR-155. We further demonstrate that miR-155 directly controls the level of TAB2, an important signal transduction molecule. Our observations suggest, therefore, that in mature human DCs, miR-155 is part of a negative feedback loop, which down-modulates inflammatory cytokine production in response to microbial stimuli.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
-
- Steinman RM, Banchereau J. Taking dendritic cells into medicine. Nature. 2007;449:419–426. - PubMed
-
- Banchereau J, Steinman RM. Dendritic cells and the control of immunity. Nature. 1998;392:245–252. - PubMed
-
- Reis e Sousa C. Dendritic cells in a mature age. Nat Rev Immunol. 2006;6:476–483. - PubMed
-
- Kawai T, Akira S. TLR signaling. Cell Death Differ. 2006;13:816–825. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

