Genome-scale analysis to the impact of gene deletion on the metabolism of E. coli: constraint-based simulation approach
- PMID: 19208166
- PMCID: PMC2648778
- DOI: 10.1186/1471-2105-10-S1-S62
Genome-scale analysis to the impact of gene deletion on the metabolism of E. coli: constraint-based simulation approach
Abstract
Background: Genome-scale models of metabolism have only been analyzed with the constraint-based modelling philosophy. Some gene deletion studies on in silico organism models at genome-scale have been made, but most of them were from the aspects of distinguishing lethal and non-lethal genes or growth rate. The impact of gene deletion on flux redistribution, the functions and characters of key genes, and the performance of different reactions in entire gene deletion still lack research.
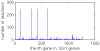
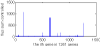
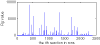
Results: Three main researches have been done into the metabolism of E. coli in gene deletion. The first work was about finding key genes and subsystems: First, by calculating the deletion impact p of whole 1261 genes, one by one, on the metabolic flux redistribution of E. coli_iAF1260, we can find that p is more detailed in describing the change of organism's metabolism. Next, we sought out 195 important (high-p) genes, and they are more than essential genes (growth rate f becomes zero if deleting). So we speculated that under some circumstances and when an important gene is deleted, a big change in the metabolic system of E. coli has taken place and E. coli may use other reaction ways to strive to live. Further, by determining the functional subsystems to which 195 key genes belong, we found that their distribution to subsystems was not even and most of them were related to just three subsystems and that all of the 8 important but not essential genes appear just in "Oxidative Phosphorylation". Our second work was about p's three characters: We analyzed the correlation between p and d (connection degree of one gene) and the correlation between p and vgene (flux sum controlled by one gene), and found that both of them are not of linear correlation, but the correlation between p and f is of highly linear correlation. The third work was about highly-affected reactions: We found 16 reactions with more than 2000 Rg value (measuring the impact that a reaction is gotten in the whole 1261 gene deletion). We speculated that highly-affected reactions involve in the metabolism of basic biomasses.
Conclusion: To sum up, these results we obtained have biological significances and our researches will shed new light on the future researches.
Figures






References
-
- Feist AM, Henry CS, Reed JL, Krummenacker M, Joyce AR, Karp PD, Broadbelt LJ, Hatzimanikatis V, Palsson BØ. A genome-scale metabolic reconstruction for Escherichia coli K-12 MG1655 that accounts for 1260 ORFs and thermodynamic information. Molecular Systems Biology. 2007;3 Art. No. 121. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

