Lin-28B transactivation is necessary for Myc-mediated let-7 repression and proliferation
- PMID: 19211792
- PMCID: PMC2651245
- DOI: 10.1073/pnas.0808300106
Lin-28B transactivation is necessary for Myc-mediated let-7 repression and proliferation
Abstract
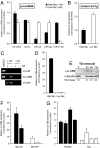
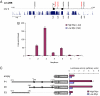
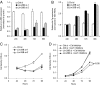
Direct control of microRNA (miRNA) expression by oncogenic and tumor suppressor networks results in frequent dysregulation of miRNAs in cancer cells and contributes to tumorigenesis. We previously demonstrated that activation of the c-Myc oncogenic transcription factor (Myc) broadly influences miRNA expression and in particular leads to widespread miRNA down-regulation. miRNA transcripts repressed by Myc include several with potent tumor suppressor activity such as miR-15a/16-1, miR-34a, and let-7 family members. In this study, we have investigated mechanisms downstream of Myc that contribute to miRNA repression. Consistent with transcriptional down-regulation, Myc activity results in the decreased abundance of multiple miRNA primary transcripts. Surprisingly, however, primary transcripts encoding several let-7 miRNAs are not reduced in the high Myc state, suggesting a posttranscriptional mechanism of repression. The Lin-28 and Lin-28B RNA binding proteins were recently demonstrated to negatively regulate let-7 biogenesis. We now show that Myc induces Lin-28B expression in multiple human and mouse tumor models. Chromatin immunoprecipitation and reporter assays reveal direct association of Myc with the Lin-28B promoter resulting in transcriptional transactivation. Moreover, we document that activation of Lin-28B is necessary and sufficient for Myc-mediated let-7 repression. Accordingly, Lin-28B loss-of-function significantly impairs Myc-dependent cellular proliferation. These findings highlight an important role for Lin-28B in Myc-driven cellular phenotypes and uncover an orchestration of transcriptional and posttranscriptional mechanisms in Myc-mediated reprogramming of miRNA expression.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
-
- Farazi TA, Juranek SA, Tuschl T. The growing catalog of small RNAs and their association with distinct Argonaute/Piwi family members. Development. 2008;135:1201–1214. - PubMed
-
- Kloosterman WP, Plasterk RH. The diverse functions of microRNAs in animal development and disease. Dev Cell. 2006;11:441–450. - PubMed
-
- Kim VN. MicroRNA biogenesis: Coordinated cropping and dicing. Nat Rev Mol Cell Biol. 2005;6:376–385. - PubMed
-
- Brennecke J, Hipfner DR, Stark A, Russell RB, Cohen SM. bantam encodes a developmentally regulated microRNA that controls cell proliferation and regulates the proapoptotic gene hid in Drosophila. Cell. 2003;113:25–36. - PubMed
-
- Reinhart BJ, et al. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature. 2000;403:901–906. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

