K63-specific deubiquitination by two JAMM/MPN+ complexes: BRISC-associated Brcc36 and proteasomal Poh1
- PMID: 19214193
- PMCID: PMC2666030
- DOI: 10.1038/emboj.2009.27
K63-specific deubiquitination by two JAMM/MPN+ complexes: BRISC-associated Brcc36 and proteasomal Poh1
Abstract
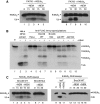
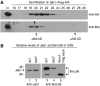
An unusual deubiquitinating (DUB) activity exists in HeLa cell extracts that is highly specific for cleaving K63-linked but not K48-linked polyubiquitin chains. The activity is insensitive to both N-ethyl-maleimide and ubiquitin aldehyde, indicating that it lacks an active site cysteine residue, and gel filtration experiments show that it resides in a high molecular weight (approximately 600 kDa) complex. Using a biochemical approach, we found that the K63-specific DUB activity co-fractionated through seven chromatographic steps with three multisubunit complexes: the 19S (PA700) portion of the 26S proteasome, the COP9 signalosome (CSN) and a novel complex that includes the JAMM/MPN+ domain-containing protein Brcc36. When we analysed the individual complexes, we found that the activity was intrinsic to PA700 and the Brcc36 isopeptidase complex (BRISC), but that the CSN-associated activity was due entirely to an interaction with Brcc36. None of the complexes cleave K6, K11, K29, K48 or alpha-linked polyubiquitin, but they do cleave K63 linkages within mixed-linkage chains. Our results suggest that specificity for K63-linked polyubiquitin is a common property of the JAMM/MPN+ family of DUBs.
Figures





Comment in
-
A set of surgical chain saws.EMBO J. 2009 Mar 18;28(6):615-6. doi: 10.1038/emboj.2009.52. EMBO J. 2009. PMID: 19295498 Free PMC article. No abstract available.
References
-
- Baboshina OV, Haas AL (1996) Novel multiubiquitin chain linkages catalyzed by the conjugating enzymes E2EPF and RAD6 are recognized by 26 S proteasome subunit 5. J Biol Chem 271: 2823–2831 - PubMed
-
- Bignell GR, Warren W, Seal S, Takahashi M, Rapley E, Barfoot R, Green H, Brown C, Biggs PJ, Lakhani SR, Jones C, Hansen J, Blair E, Hofmann B, Siebert R, Turner G, Evans DG, Schrander-Stumpel C, Beemer FA, van Den Ouweland A et al. (2000) Identification of the familial cylindromatosis tumour-suppressor gene. Nat Genet 25: 160–165 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

