A novel chromosome 19p13.12 deletion in a child with multiple congenital anomalies
- PMID: 19215039
- PMCID: PMC2872113
- DOI: 10.1002/ajmg.a.32691
A novel chromosome 19p13.12 deletion in a child with multiple congenital anomalies
Abstract
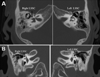
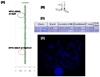
We describe a patient with multiple congenital anomalies including deafness, lacrimal duct stenosis, strabismus, bilateral cervical sinuses, congenital cardiac defects, hypoplasia of the corpus callosum, and hypoplasia of the cerebellar vermis. Mutation analysis of EYA1, SIX1, and SIX5, genes that underlie otofaciocervical and/or branchio-oto-renal syndrome, was negative. Pathologic diagnosis of the excised cervical sinus tracts was revised on re-examination to heterotopic salivary gland tissue. Using high resolution chromosomal microarray analysis, we identified a novel 2.52 Mb deletion at 19p13.12, which was confirmed by fluorescent in situ hybridization and demonstrated to be a de novo mutation by testing of the parents. Overall, deletions of chromosome 19p13 are rare.
2009 Wiley-Liss, Inc.
Figures




References
-
- Ain Q, Nazli S, Riazuddin S, Jaleel AU, Riazuddin SA, Zafar AU, Khan SN, Husnain T, Griffith AJ, Ahmed ZM, Friedman TB. The autosomal recessive nonsyndromic deafness locus DFNB72 is located on chromosome 19p13.3. Hum Genet. 2007;122:445–450. - PubMed
-
- Archer HL, Gupta S, Enoch S, Thompson P, Rowbottom A, Chua I, Warren S, Johnson D, Ledbetter DH, Lese-Martin C, Williams P, Pilz DT. Distinct phenotype associated with a cryptic subtelomeric deletion of 19p13.3-pter. Am J Med Genet Part A. 2005;136A:38–44. - PubMed
-
- Bergstrom L, Hemenway WG, Downs MP. A high risk registry to find congenital deafness. Otolaryngol Clin North Am. 1971;4:369–399. - PubMed
-
- Bitner-Glindzicz M. Hereditary deafness and phenotyping in humans. Br Med Bull. 2002;63:73–94. - PubMed
-
- Bonsch D, Schmidt CM, Scheer P, Bohlender J, Neumann C, am Zehnhoff-Dinnesen A, Deufel T. [A new locus for an autosomal dominant, non-syndromic hearing impairment (DFNA57) located on chromosome 19p13.2 and overlapping with DFNB15] HNO. 2008;56:177–182. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

