The Phycodnaviridae: the story of how tiny giants rule the world
- PMID: 19216434
- PMCID: PMC2908299
- DOI: 10.1007/978-3-540-68618-7_1
The Phycodnaviridae: the story of how tiny giants rule the world
Abstract
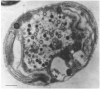
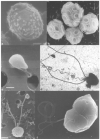
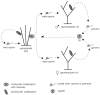
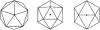
The family Phycodnaviridae encompasses a diverse and rapidly expanding collection of large icosahedral, dsDNA viruses that infect algae. These lytic and lysogenic viruses have genomes ranging from 160 to 560 kb. The family consists of six genera based initially on host range and supported by sequence comparisons. The family is monophyletic with branches for each genus, but the phycodnaviruses have evolutionary roots that connect them with several other families of large DNA viruses, referred to as the nucleocytoplasmic large DNA viruses (NCLDV). The phycodnaviruses have diverse genome structures, some with large regions of noncoding sequence and others with regions of ssDNA. The genomes of members in three genera in the Phycodnaviridae have been sequenced. The genome analyses have revealed more than 1000 unique genes, with only 14 homologous genes in common among the three genera of phycodnaviruses sequenced to date. Thus, their gene diversity far exceeds the number of so-called core genes. Not much is known about the replication of these viruses, but the consequences of these infections on phytoplankton have global affects, including influencing geochemical cycling and weather patterns.
Figures

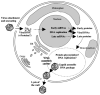

References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

