Recognition and processing of a new repertoire of DNA substrates by human 3-methyladenine DNA glycosylase (AAG)
- PMID: 19219989
- PMCID: PMC2883313
- DOI: 10.1021/bi8018898
Recognition and processing of a new repertoire of DNA substrates by human 3-methyladenine DNA glycosylase (AAG)
Abstract
The human 3-methyladenine DNA glycosylase (AAG) recognizes and excises a broad range of purines damaged by alkylation and oxidative damage, including 3-methyladenine, 7-methylguanine, hypoxanthine (Hx), and 1,N(6)-ethenoadenine (epsilonA). The crystal structures of AAG bound to epsilonA have provided insights into the structural basis for substrate recognition, base excision, and exclusion of normal purines and pyrimidines from its substrate recognition pocket. In this study, we explore the substrate specificity of full-length and truncated Delta80AAG on a library of oligonucleotides containing structurally diverse base modifications. Substrate binding and base excision kinetics of AAG with 13 damaged oligonucleotides were examined. We found that AAG bound to a wide variety of purine and pyrimidine lesions but excised only a few of them. Single-turnover excision kinetics showed that in addition to the well-known epsilonA and Hx substrates, 1-methylguanine (m1G) was also excised efficiently by AAG. Thus, along with epsilonA and ethanoadenine (EA), m1G is another substrate that is shared between AAG and the direct repair protein AlkB. In addition, we found that both the full-length and truncated AAG excised 1,N(2)-ethenoguanine (1,N(2)-epsilonG), albeit weakly, from duplex DNA. Uracil was excised from both single- and double-stranded DNA, but only by full-length AAG, indicating that the N-terminus of AAG may influence glycosylase activity for some substrates. Although AAG has been primarily shown to act on double-stranded DNA, AAG excised both epsilonA and Hx from single-stranded DNA, suggesting the possible significance of repair of these frequent lesions in single-stranded DNA transiently generated during replication and transcription.
Figures
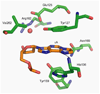
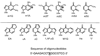

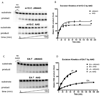
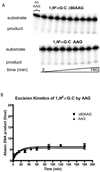
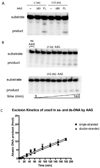
 ) full-length AAG.
) full-length AAG.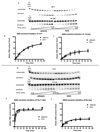
Similar articles
-
Effects of substrate specificity on initiating the base excision repair of N-methylpurines by variant human 3-methyladenine DNA glycosylases.Chem Res Toxicol. 2005 Jan;18(1):87-94. doi: 10.1021/tx049822q. Chem Res Toxicol. 2005. PMID: 15651853
-
Human alkyladenine DNA glycosylase uses acid-base catalysis for selective excision of damaged purines.Biochemistry. 2003 Oct 28;42(42):12418-29. doi: 10.1021/bi035177v. Biochemistry. 2003. PMID: 14567703
-
Base excision and DNA binding activities of human alkyladenine DNA glycosylase are sensitive to the base paired with a lesion.J Biol Chem. 2001 Apr 20;276(16):13379-87. doi: 10.1074/jbc.M010641200. Epub 2001 Jan 22. J Biol Chem. 2001. PMID: 11278716
-
DNA glycosylase recognition and catalysis.Curr Opin Struct Biol. 2004 Feb;14(1):43-9. doi: 10.1016/j.sbi.2004.01.003. Curr Opin Struct Biol. 2004. PMID: 15102448 Review.
-
Structural studies of human alkyladenine glycosylase and E. coli 3-methyladenine glycosylase.Mutat Res. 2000 Aug 30;460(3-4):201-10. doi: 10.1016/s0921-8777(00)00027-6. Mutat Res. 2000. PMID: 10946229 Review.
Cited by
-
Unraveling radiation resistance strategies in two bacterial strains from the high background radiation area of Chavara-Neendakara: A comprehensive whole genome analysis.PLoS One. 2024 Jun 10;19(6):e0304810. doi: 10.1371/journal.pone.0304810. eCollection 2024. PLoS One. 2024. PMID: 38857267 Free PMC article.
-
Alkyladenine DNA glycosylase (AAG) localizes to mitochondria and interacts with mitochondrial single-stranded binding protein (mtSSB).DNA Repair (Amst). 2013 Mar 1;12(3):177-87. doi: 10.1016/j.dnarep.2012.11.009. Epub 2013 Jan 3. DNA Repair (Amst). 2013. PMID: 23290262 Free PMC article.
-
Mitochondrial DNA damage and its consequences for mitochondrial gene expression.Biochim Biophys Acta. 2012 Sep-Oct;1819(9-10):979-91. doi: 10.1016/j.bbagrm.2012.06.002. Epub 2012 Jun 19. Biochim Biophys Acta. 2012. PMID: 22728831 Free PMC article. Review.
-
N-methylpurine DNA glycosylase and OGG1 DNA repair activities: opposite associations with lung cancer risk.J Natl Cancer Inst. 2012 Nov 21;104(22):1765-9. doi: 10.1093/jnci/djs445. Epub 2012 Oct 27. J Natl Cancer Inst. 2012. PMID: 23104324 Free PMC article.
-
The Mbd4 DNA glycosylase protects mice from inflammation-driven colon cancer and tissue injury.Oncotarget. 2016 May 10;7(19):28624-36. doi: 10.18632/oncotarget.8721. Oncotarget. 2016. PMID: 27086921 Free PMC article.
References
-
- Friedberg EC, Walker GC, Siede W, Wood RD, Schultz RA, Ellenberger T. DNA Repair and Mutagenesis. 2nd edition. Washington DC: ASM press; 2006.
-
- Gallagher PE, Brent TP. Partial purification and characterization of 3-methyladenine-DNA glycosylase from human placenta. Biochemistry. 1982;21:6404–6409. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- R37 CA080024/CA/NCI NIH HHS/United States
- R01 CA055042/CA/NCI NIH HHS/United States
- ES02109/ES/NIEHS NIH HHS/United States
- CA26731/CA/NCI NIH HHS/United States
- CA55042/CA/NCI NIH HHS/United States
- T32 ES007020/ES/NIEHS NIH HHS/United States
- ES0535/ES/NIEHS NIH HHS/United States
- P30 ES002109/ES/NIEHS NIH HHS/United States
- P01 ES005355/ES/NIEHS NIH HHS/United States
- T32-ES007020/ES/NIEHS NIH HHS/United States
- R01 CA080024/CA/NCI NIH HHS/United States
- R01 CA086061/CA/NCI NIH HHS/United States
- R01 CA075576/CA/NCI NIH HHS/United States
- P01 CA026731/CA/NCI NIH HHS/United States
- CA80024/CA/NCI NIH HHS/United States
- CA75576/CA/NCI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources

