Indexing TNF-alpha gene expression using a gene-targeted reporter cell line
- PMID: 19220876
- PMCID: PMC2657777
- DOI: 10.1186/1741-7007-7-8
Indexing TNF-alpha gene expression using a gene-targeted reporter cell line
Abstract
Background: Current cell-based drug screening technologies utilize randomly integrated reporter genes to index transcriptional activity of an endogenous gene of interest. In this context, reporter expression is controlled by known genetic elements that may only partially capture gene regulation and by unknown features of chromatin specific to the integration site. As an alternative technology, we applied highly efficient gene-targeting with recombinant adeno-associated virus to precisely integrate a luciferase reporter gene into exon 1 of the HeLa cell tumor necrosis factor-alpha (TNF-alpha) gene. Drugs known to induce TNF-alpha expression were then used to compare the authenticity of gene-targeted and randomly integrated transcriptional reporters.
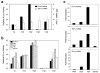
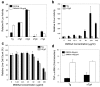
Results: TNF-alpha-targeted reporter activity reflected endogenous TNF-alpha mRNA expression, whereas randomly integrated TNF-alpha reporter lines gave variable expression in response to transcriptional and epigenetic regulators. 5,6-Dimethylxanthenone-4-acetic acid (DMXAA), currently used in cancer clinical trials to induce TNF-alpha gene transcription, was only effective at inducing reporter expression from TNF-alpha gene-targeted cells.
Conclusion: We conclude that gene-targeted reporter cell lines provide predictive indexing of gene transcription for drug discovery.
Figures


References
-
- Carey M, Smale ST. Transcriptional regulation in eukaryotes: concept, strategies, and techniques. Cold Spring Harbor, NY: CHS Laboratory Press; 2000.
-
- Kain SR, Ganguly S. Overview of genetic reporter systems. Curr Protoc Mol Biol. 2001;Chapter 9:Unit 9.6. - PubMed
-
- Swevers L, Kravariti L, Ciolfi S, Xenou-Kokoletsi M, Ragoussis N, Smagghe G, Nakagawa Y, Mazomenos B, Iatrou K. A cell-based high-throughput screening system for detecting ecdysteroid agonists and antagonists in plant extracts and libraries of synthetic compounds. Faseb J. 2004;18:134–136. - PubMed
-
- Tang Y, Luo J, Fleming CR, Kong Y, Olini GC, Jr, Wildey MJ, Cavender DE, Demarest KT. Development of a sensitive and HTS-compatible reporter gene assay for functional analysis of human adenosine A2a receptors in CHO-K1 cells. Assay Drug Dev Technol. 2004;2:281–289. doi: 10.1089/1540658041410650. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

