Monomethylation of histone H4-lysine 20 is involved in chromosome structure and stability and is essential for mouse development
- PMID: 19223465
- PMCID: PMC2663305
- DOI: 10.1128/MCB.01768-08
Monomethylation of histone H4-lysine 20 is involved in chromosome structure and stability and is essential for mouse development
Abstract
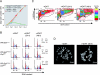
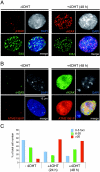
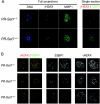
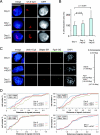
PR-Set7/Set8/KMT5A is the sole enzyme known to catalyze monomethylation of histone H4 lysine 20 (H4K20) and is present only in multicellular organisms that compact a large fraction of their DNA. We found that mouse embryos that are homozygous null mutants for the gene PR-Set7 display early embryonic lethality prior to the eight-cell stage. Death was due to the absence of PR-Set7 catalytic activity, since microinjection of the wild type, but not a catalytically inactive version, into two-cell embryos rescued the phenotype. A lack of PR-Set7 activity resulted not only in depletion of H4K20me1 but also in reduced levels of the H4K20me2/3 marks catalyzed by the Suv4-20h1/h2 enzymes, implying that H4K20me1 may be essential for the function of these enzymes to ensure the dimethylated and trimethylated states. Embryonic stem cells that were inducibly deleted for PR-Set7 passed through an initial G(2)/M phase, but the progeny were defective at the subsequent S and G(2)/M phases, exhibiting a delay in their cell cycle, accumulation at G(2)/M, massive DNA damage, and improper mitotic chromosome condensation. Cell cycle analysis after synchronization indicated that the defects were a consequence of decreased H4K20me1 due to the absence of PR-Set7. Most importantly, the lack of H4K20me1 also resulted in defects in chromosome condensation in interphase nuclei. These results demonstrate the critical role of H4K20 monomethylation in mammals in a developmental context.
Figures







References
-
- Barski, A., S. Cuddapah, K. Cui, T. Y. Roh, D. E. Schones, Z. Wang, G. Wei, I. Chepelev, and K. Zhao. 2007. High-resolution profiling of histone methylations in the human genome. Cell 129823-837. - PubMed
-
- Dorigo, B., T. Schalch, K. Bystricky, and T. J. Richmond. 2003. Chromatin fiber folding: requirement for the histone H4 N-terminal tail. J. Mol. Biol. 32785-96. - PubMed
-
- Fraga, M. F., E. Ballestar, A. Villar-Garea, M. Boix-Chornet, J. Espada, G. Schotta, T. Bonaldi, C. Haydon, S. Ropero, K. Petrie, N. G. Iyer, A. Perez-Rosado, E. Calvo, J. A. Lopez, A. Cano, M. J. Calasanz, D. Colomer, M. A. Piris, N. Ahn, A. Imhof, C. Caldas, T. Jenuwein, and M. Esteller. 2005. Loss of acetylation at Lys16 and trimethylation at Lys20 of histone H4 is a common hallmark of human cancer. Nat. Genet. 37391-400. - PubMed
-
- Hayashi, S., P. Lewis, L. Pevny, and A. P. McMahon. 2002. Efficient gene modulation in mouse epiblast using a Sox2Cre transgenic mouse strain. Mech. Dev. 119(Suppl. 1)S97-S101. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
